Abstract
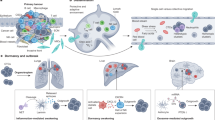
Transposable elements (TEs) represent almost half of the human genome. Historically deemed ‘junk DNA’, recent technological advancements have stimulated a wave of research into the functional impact of TEs on gene-regulatory networks in evolution and development, as well as in diseases including cancer. The genetic and epigenetic evolution of cancer involves the exploitation of TEs, whereby TEs contribute directly to cancer-specific gene activities. This Review provides a perspective on the role of TEs in cancer as being a ‘double-edged sword’, both promoting cancer evolution and representing a vulnerability that could be exploited in cancer therapy. We discuss how TEs affect transcriptome regulation and other cellular processes in cancer. We highlight the potential of TEs as therapeutic targets for cancer. We also summarize technical hurdles in the characterization of TEs with genomic assays. Last, we outline open questions and exciting future research avenues.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hanahan, D. Hallmarks of cancer: new dimensions. Cancer Discov. 12, 31–46 (2022).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Black, J. R. M. & McGranahan, N. Genetic and non-genetic clonal diversity in cancer evolution. Nat. Rev. Cancer 21, 379–392 (2021).
Davila, M. L. & Brentjens, R. J. CAR T cell therapy: looking back and looking forward. Nat. Cancer 3, 1418–1419 (2022).
Cordaux, R. & Batzer, M. A. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 10, 691–703 (2009).
Feschotte, C. & Pritham, E. J. DNA transposons and the evolution of eukaryotic genomes. Annu. Rev. Genet. 41, 331–368 (2007).
Hartl, D. L., Lozovskaya, E. R. & Lawrence, J. G. Nonautonomous transposable elements in prokaryotes and eukaryotes. Genetica 86, 47–53 (1992).
Sun, C., Feschotte, C., Wu, Z. & Mueller, R. L. DNA transposons have colonized the genome of the giant virus Pandoravirus salinus. BMC Biol. 13, 38 (2015).
Wicker, T. et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 8, 973–982 (2007).
Kazazian, H. H. & Moran, J. V. Mobile DNA in health and disease. N. Engl. J. Med. 377, 361–370 (2017).
Burns, K. H. Transposable elements in cancer. Nat. Rev. Cancer 17, 415–424 (2017).
Modzelewski, A. J., Gan Chong, J., Wang, T. & He, L. Mammalian genome innovation through transposon domestication. Nat. Cell Biol. 24, 1332–1340 (2022).
Fueyo, R. Roles of transposable elements in the regulation of mammalian transcription. Nat. Rev. Mol. Cell Biol. 23, 481–497 (2022).
Lawson, H. A., Liang, Y. & Wang, T. Transposable elements in mammalian chromatin organization. Nat. Rev. Genet. 24, 712–723 (2023).
Hancks, D. C. & Kazazian, H. H. Roles for retrotransposon insertions in human disease. Mob. DNA 7, 9 (2016).
Payer, L. M. et al. Structural variants caused by Alu insertions are associated with risks for many human diseases. Proc. Natl Acad. Sci. USA 114, E3984–E3992 (2017).
Beck, C. R., Garcia-Perez, J. L., Badge, R. M. & Moran, J. V. LINE-1 elements in structural variation and disease. Annu. Rev. Genom. Hum. Genet. 12, 187–215 (2011).
Kazazian, H. H. et al. Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature 332, 164–166 (1988).
Miki, Y. et al. Disruption of the APC gene by a retrotransposal insertion of L1 sequence in a colon cancer. Cancer Res. 52, 643–645 (1992).
Scott, E. C. et al. A hot L1 retrotransposon evades somatic repression and initiates human colorectal cancer. Genome Res. 26, 745–755 (2016).
Cajuso, T. et al. Retrotransposon insertions can initiate colorectal cancer and are associated with poor survival. Nat. Commun. 10, 4022 (2019).
Nam, C. H. et al. Widespread somatic L1 retrotransposition in normal colorectal epithelium. Nature 617, 540–547 (2023).
Doucet-O’Hare, T. T. et al. LINE-1 expression and retrotransposition in Barrett’s esophagus and esophageal carcinoma. Proc. Natl Acad. Sci. USA 112, E4894–E4900 (2015).
Katz-Summercorn, A. C. et al. Multi-omic cross-sectional cohort study of pre-malignant Barrett’s esophagus reveals early structural variation and retrotransposon activity. Nat. Commun. 13, 1407 (2022).
Shukla, R. et al. Endogenous retrotransposition activates oncogenic pathways in hepatocellular carcinoma. Cell 153, 101–111 (2013).
Rodriguez-Martin, B. et al. Pan-cancer analysis of whole genomes identifies driver rearrangements promoted by LINE-1 retrotransposition. Nat. Genet. 52, 306–319 (2020).
Scott, E. & Devine, S. The role of somatic L1 retrotransposition in human cancers. Viruses 9, 131 (2017).
Ewing, A. D. et al. Widespread somatic L1 retrotransposition occurs early during gastrointestinal cancer evolution. Genome Res. 25, 1536–1545 (2015).
Ewing, A. D. et al. Nanopore sequencing enables comprehensive transposable element epigenomic profiling. Mol. Cell 80, 915–928.e5 (2020).
Ardeljan, D. et al. Cell fitness screens reveal a conflict between LINE-1 retrotransposition and DNA replication. Nat. Struct. Mol. Biol. 27, 168–178 (2020).
Belgnaoui, S. M., Gosden, R. G., Semmes, O. J. & Haoudi, A. Human LINE-1 retrotransposon induces DNA damage and apoptosis in cancer cells. Cancer Cell Int. 6, 13 (2006).
Gu, Z. et al. Silencing of LINE-1 retrotransposons is a selective dependency of myeloid leukemia. Nat. Genet. 53, 672–682 (2021).
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Kellner, M. & Makałowski, W. Transposable elements significantly contributed to the core promoters in the human genome. Sci. China Life Sci. 62, 489–497 (2019).
Huda, A., Bowen, N. J., Conley, A. B. & Jordan, I. K. Epigenetic regulation of transposable element derived human gene promoters. Gene 475, 39–48 (2011).
Sundaram, V. & Wysocka, J. Transposable elements as a potent source of diverse cis-regulatory sequences in mammalian genomes. Phil. Trans. R. Soc. B 375, 20190347 (2020).
Lynch-Sutherland, C. F., Chatterjee, A., Stockwell, P. A., Eccles, M. R. & Macaulay, E. C. Reawakening the developmental origins of cancer through transposable elements. Front. Oncol. 10, 468 (2020).
Santoni, F. A., Guerra, J. & Luban, J. HERV-H RNA is abundant in human embryonic stem cells and a precise marker for pluripotency. Retrovirology 9, 111 (2012).
Pontis, J. et al. Hominoid-specific transposable elements and KZFPs facilitate human embryonic genome activation and control transcription in naive human ESCs. Cell Stem Cell 24, 724–735.e5 (2019).
Macfarlan, T. S. et al. Embryonic stem cell potency fluctuates with endogenous retrovirus activity. Nature 487, 57–63 (2012).
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat. Genet. 42, 631–634 (2010).
FANTOM Consortium et al. Deep transcriptome profiling of mammalian stem cells supports a regulatory role for retrotransposons in pluripotency maintenance. Nat. Genet. 46, 558–566 (2014).
Lu, X. et al. The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nat. Struct. Mol. Biol. 21, 423–425 (2014).
Wang, J. et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 516, 405–409 (2014).
Modzelewski, A. J. et al. A mouse-specific retrotransposon drives a conserved Cdk2ap1 isoform essential for development. Cell 184, 5541–5558.e22 (2021).
Shademan, M. et al. Promoter methylation, transcription, and retrotransposition of LINE-1 in colorectal adenomas and adenocarcinomas. Cancer Cell Int. 20, 426 (2020).
Kong, Y. et al. Transposable element expression in tumors is associated with immune infiltration and increased antigenicity. Nat. Commun. 10, 5228 (2019). This study shows that transposable elements are associated with increased sensitivity of tumours to the immune system, possibly by augmenting the neoantigen repository and increasing abundance of double-stranded RNA.
Iouranova, A. et al. KRAB zinc finger protein ZNF676 controls the transcriptional influence of LTR12-related endogenous retrovirus sequences. Mob. DNA 13, 4 (2022).
Ecco, G. et al. Transposable elements and their KRAB-ZFP controllers regulate gene expression in adult tissues. Dev. Cell 36, 611–623 (2016).
Wylie, A. et al. p53 genes function to restrain mobile elements. Genes. Dev. 30, 64–77 (2016).
Tiwari, B. et al. p53 directly represses human LINE1 transposons. Genes. Dev. 34, 1439–1451 (2020).
Yu, C. et al. ARID1A loss derepresses a group of human endogenous retrovirus-H loci to modulate BRD4-dependent transcription. Nat. Commun. 13, 3501 (2022).
Gainetdinov, I., Skvortsova, Y., Kondratieva, S., Funikov, S. & Azhikina, T. Two modes of targeting transposable elements by piRNA pathway in human testis. RNA 23, 1614–1625 (2017).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007).
Gunawardane, L. S. et al. A slicer-mediated mechanism for repeat-associated siRNA 5' end formation in Drosophila. Science 315, 1587–1590 (2007).
Wolff, E. M. et al. Hypomethylation of a LINE-1 promoter activates an alternate transcript of the MET oncogene in bladders with cancer. PLoS Genet. 6, e1000917 (2010).
Edginton-White, B. et al. Global long terminal repeat activation participates in establishing the unique gene expression programme of classical Hodgkin lymphoma. Leukemia 33, 1463–1474 (2019).
Feinberg, A. P. & Vogelstein, B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature 301, 89–92 (1983).
Nigumann, P., Redik, K., Mätlik, K. & Speek, M. Many human genes are transcribed from the antisense promoter of L1 retrotransposon. Genomics 79, 628–634 (2002).
Grundy, E. E., Diab, N. & Chiappinelli, K. B. Transposable element regulation and expression in cancer. FEBS J. 289, 1160–1179 (2022).
Zhao, Y. et al. Transposon-triggered innate immune response confers cancer resistance to the blind mole rat. Nat. Immunol. 22, 1219–1230 (2021).
Cherkasova, E. et al. Detection of an immunogenic HERV-E envelope with selective expression in clear cell kidney cancer. Cancer Res. 76, 2177–2185 (2016).
Deblois, G. et al. Epigenetic switch-induced viral mimicry evasion in chemotherapy-resistant breast cancer. Cancer Discov. 10, 1312–1329 (2020).
Topham, J. T. et al. Endogenous retrovirus transcript levels are associated with immunogenic signatures in multiple metastatic cancer types. Mol. Cancer Ther. 19, 1889–1897 (2020).
Jang, H. S. et al. Transposable elements drive widespread expression of oncogenes in human cancers. Nat. Genet. 51, 611–617 (2019). This study demonstrates that transposable elements can function as cryptic oncogenic promoters that drive oncogene overexpression and facilitate tumorigenesis.
Shah, N. M. et al. Pan-cancer analysis identifies tumor-specific antigens derived from transposable elements. Nat. Genet. 55, 631–639 (2023). This study comprehensively profiles the expression landscape of tumour-specific transposable-element–gene chimeric transcripts in a pan-cancer manner, demonstrating the potential for off-the-shelf transposable-element-derived vaccines and other immunotherapy applications targeting transposable elements.
Sato, S. et al. LINE-1 ORF1p as a candidate biomarker in high grade serous ovarian carcinoma. Sci. Rep. 13, 1537 (2023).
Rodić, N. et al. Long interspersed element-1 protein expression is a hallmark of many human cancers. Am. J. Pathol. 184, 1280–1286 (2014).
Rooney, M. S., Shukla, S. A., Wu, C. J., Getz, G. & Hacohen, N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell 160, 48–61 (2015).
Smith, C. C. et al. Endogenous retroviral signatures predict immunotherapy response in clear cell renal cell carcinoma. J. Clin. Invest. 128, 4804–4820 (2018). This study shows that the expression of transposable elements can predict immunotherapy response in clear cell renal cell carcinoma, indirectly indicating that the expression of transposable elements can sensitize tumours to the immune system.
Babaian, A. & Mager, D. L. Endogenous retroviral promoter exaptation in human cancer. Mob. DNA 7, 24 (2016).
Attig, J. et al. LTR retroelement expansion of the human cancer transcriptome and immunopeptidome revealed by de novo transcript assembly. Genome Res. 29, 1578–1590 (2019).
Chuong, E. B., Rumi, M. A. K., Soares, M. J. & Baker, J. C. Endogenous retroviruses function as species-specific enhancer elements in the placenta. Nat. Genet. 45, 325–329 (2013).
Ye, M. et al. Specific subfamilies of transposable elements contribute to different domains of T lymphocyte enhancers. Proc. Natl Acad. Sci. USA 117, 7905–7916 (2020).
Liang, L. et al. Complementary Alu sequences mediate enhancer–promoter selectivity. Nature 619, 868–875 (2023).
Fuentes, D. R., Swigut, T. & Wysocka, J. Systematic perturbation of retroviral LTRs reveals widespread long-range effects on human gene regulation. eLife 7, e35989 (2018).
Karttunen, K. et al. Transposable elements as tissue-specific enhancers in cancers of endodermal lineage. Nat. Commun. 14, 5313 (2023).
Deniz, Ö. et al. Endogenous retroviruses are a source of enhancers with oncogenic potential in acute myeloid leukaemia. Nat. Commun. 11, 3506 (2020). This study validates the activity of transposable-element-derived enhancers in the context of cancer and shows the oncogenic function of a subset of them.
Grillo, G. et al. Transposable elements are co-opted as oncogenic regulatory elements by lineage-specific transcription factors in prostate cancer. Cancer Discov. 13, 2470–2487 (2023).
Schmid, C. D. & Bucher, P. MER41 repeat sequences contain inducible STAT1 binding sites. PLoS One 5, e11425 (2010).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science 351, 1083–1087 (2016).
Ito, J. et al. Endogenous retroviruses drive KRAB zinc-finger protein family expression for tumor suppression. Sci. Adv. 6, eabc3020 (2020).
Belancio, V. P., Hedges, D. J. & Deininger, P. LINE-1 RNA splicing and influences on mammalian gene expression. Nucleic Acids Res. 34, 1512–1521 (2006).
Lev-Maor, G. et al. Intronic Alus influence alternative splicing. PLoS Genet. 4, e1000204 (2008).
Clayton, E. A. et al. An atlas of transposable element-derived alternative splicing in cancer. Phil. Trans. R. Soc. B 375, 20190342 (2020).
Kim, W. R. et al. Integration of TE induces cancer specific alternative splicing events. Int. J. Mol. Sci. 23, 10918 (2022).
Kahles, A. et al. Comprehensive analysis of alternative splicing across tumors from 8,705 patients. Cancer Cell 34, 211–224.e6 (2018).
Burbage, M. et al. Epigenetically controlled tumor antigens derived from splice junctions between exons and transposable elements. Sci. Immunol. 8, eabm6360 (2023).
Merlotti, A. et al. Noncanonical splicing junctions between exons and transposable elements represent a source of immunogenic recurrent neo-antigens in patients with lung cancer. Sci. Immunol. 8, eabm6359 (2023).
Zhang, Y., Qian, J., Gu, C. & Yang, Y. Alternative splicing and cancer: a systematic review. Signal. Transduct. Target. Ther. 6, 78 (2021).
Bonnal, S. C., López-Oreja, I. & Valcárcel, J. Roles and mechanisms of alternative splicing in cancer—implications for care. Nat. Rev. Clin. Oncol. 17, 457–474 (2020).
Plaisance, I. et al. A transposable element into the human long noncoding RNA CARMEN is a switch for cardiac precursor cell specification. Cardiovasc. Res. 119, 1361–1376 (2023).
Jones, P. A., Ohtani, H., Chakravarthy, A. & De Carvalho, D. D. Epigenetic therapy in immune-oncology. Nat. Rev. Cancer 19, 151–161 (2019).
Mehdipour, P. et al. Epigenetic therapy induces transcription of inverted SINEs and ADAR1 dependency. Nature 588, 169–173 (2020).
Chen, Y. G. & Hur, S. Cellular origins of dsRNA, their recognition and consequences. Nat. Rev. Mol. Cell Biol. 23, 286–301 (2022).
Elbarbary, R. A. & Maquat, L. E. Distinct mechanisms obviate the potentially toxic effects of inverted-repeat Alu elements on cellular RNA metabolism. Nat. Struct. Mol. Biol. 24, 496–498 (2017).
Heinrich, M. J. et al. Endogenous double-stranded Alu RNA elements stimulate IFN-responses in relapsing remitting multiple sclerosis. J. Autoimmun. 100, 40–51 (2019).
Wu, Q. et al. PRMT inhibition induces a viral mimicry response in triple-negative breast cancer. Nat. Chem. Biol. 18, 821–830 (2022).
Wilson, K. D. et al. Endogenous retrovirus-derived lncRNA BANCR promotes cardiomyocyte migration in humans and non-human primates. Dev. Cell 54, 694–709.e9 (2020).
Jin, X. et al. The endogenous retrovirus-derived long noncoding RNA TROJAN promotes triple-negative breast cancer progression via ZMYND8 degradation. Sci. Adv. 5, eaat9820 (2019).
Noer, J. B., Hørsdal, O. K., Xiang, X., Luo, Y. & Regenberg, B. Extrachromosomal circular DNA in cancer: history, current knowledge, and methods. Trends Genet. 38, 766–781 (2022).
Wang, T., Zhang, H., Zhou, Y. & Shi, J. Extrachromosomal circular DNA: a new potential role in cancer progression. J. Transl. Med. 19, 257 (2021).
Wang, Y. et al. eccDNAs are apoptotic products with high innate immunostimulatory activity. Nature 599, 308–314 (2021).
Møller, H. D. et al. Formation of extrachromosomal circular DNA from long terminal repeats of retrotransposons in Saccharomyces cerevisiae. G3 6, 453–462 (2016).
Yang, F. et al. Retrotransposons hijack alt-EJ for DNA replication and eccDNA biogenesis. Nature 620, 218–225 (2023).
De Cecco, M. et al. L1 drives IFN in senescent cells and promotes age-associated inflammation. Nature 566, 73–78 (2019).
Simon, M. et al. LINE1 derepression in aged wild-type and SIRT6-deficient mice drives inflammation. Cell Metab. 29, 871–885.e5 (2019).
Serafino, A. et al. The activation of human endogenous retrovirus K (HERV-K) is implicated in melanoma cell malignant transformation. Exp. Cell Res. 315, 849–862 (2009).
Li, M. et al. Downregulation of human endogenous retrovirus type K (HERV-K) viral env RNA in pancreatic cancer cells decreases cell proliferation and tumor growth. Clin. Cancer Res. 23, 5892–5911 (2017).
Zhou, F. et al. Activation of HERV-K Env protein is essential for tumorigenesis and metastasis of breast cancer cells. Oncotarget 7, 84093–84117 (2016).
Chen, T. et al. The viral oncogene Np9 acts as a critical molecular switch for co-activating β-catenin, ERK, Akt and Notch1 and promoting the growth of human leukemia stem/progenitor cells. Leukemia 27, 1469–1478 (2013).
Lamprecht, B. et al. Derepression of an endogenous long terminal repeat activates the CSF1R proto-oncogene in human lymphoma. Nat. Med. 16, 571–579 (2010).
Babaian, A. et al. Onco-exaptation of an endogenous retroviral LTR drives IRF5 expression in Hodgkin lymphoma. Oncogene 35, 2542–2546 (2016).
Scarfò, I. et al. Identification of a new subclass of ALK-negative ALCL expressing aberrant levels of ERBB4 transcripts. Blood 127, 221–232 (2016).
Wiesner, T. et al. Alternative transcription initiation leads to expression of a novel ALK isoform in cancer. Nature 526, 453–457 (2015).
Lock, F. E. et al. Distinct isoform of FABP7 revealed by screening for retroelement-activated genes in diffuse large B-cell lymphoma. Proc. Natl Acad. Sci. USA 111, E3534–E3543 (2014).
Simó-Riudalbas, L. et al. Transposon-activated POU5F1B promotes colorectal cancer growth and metastasis. Nat. Commun. 13, 4913 (2022).
Lock, F. E. et al. A novel isoform of IL-33 revealed by screening for transposable element promoted genes in human colorectal cancer. PLOS One 12, e0180659 (2017).
Grimmett, E. et al. Cancer vaccines: past, present and future; a review article. Discov. Oncol. 13, 31 (2022).
Kubli, S. P., Berger, T., Araujo, D. V., Siu, L. L. & Mak, T. W. Beyond immune checkpoint blockade: emerging immunological strategies. Nat. Rev. Drug. Discov. 20, 899–919 (2021).
Lin, M. J. et al. Cancer vaccines: the next immunotherapy frontier. Nat. Cancer 3, 911–926 (2022).
Schumacher, T. N. & Schreiber, R. D. Neoantigens in cancer immunotherapy. Science 348, 69–74 (2015).
Dougan, M., Dranoff, G. & Dougan, S. K. Cancer immunotherapy: beyond checkpoint blockade. JACC CardioOncol. 4, 563–578 (2019).
Laumont, C. M. et al. Global proteogenomic analysis of human MHC class I-associated peptides derived from non-canonical reading frames. Nat. Commun. 7, 10238 (2016).
Laumont, C. M. et al. Noncoding regions are the main source of targetable tumor-specific antigens. Sci. Transl. Med. 10, eaau5516 (2018).
Zhao, Q. et al. Proteogenomics uncovers a vast repertoire of shared tumor-specific antigens in ovarian cancer. Cancer Immunol. Res. 8, 544–555 (2020).
Takahashi, Y. et al. Regression of human kidney cancer following allogeneic stem cell transplantation is associated with recognition of an HERV-E antigen by T cells. J. Clin. Invest. 118, 1099–1109 (2008).
Krishnamurthy, J. et al. Genetic engineering of T cells to target HERV-K, an ancient retrovirus on melanoma. Clin. Cancer Res. 21, 3241–3251 (2015). This study provides an example of chimeric antigen receptor T cells engineered to target transposable elements leading to successful tumor clearance in a melanoma mouse model.
Bonaventura, P., Alain, V., Qing, W., Christophe, C. & Stéphane, D. Identification of shared tumor epitopes from endogenous retroviruses inducing high-avidity cytotoxic T cells for cancer immunotherapy. Sci. Adv. 8, eabj3671 (2022).
Ng, K. W. et al. Antibodies against endogenous retroviruses promote lung cancer immunotherapy. Nature 616, 563–573 (2023). This study showcases an example of a successful antibody-based immunotherapy treatment targeting transposable elements in a lung cancer mouse model.
Bonté, P.-E. et al. Single-cell RNA-seq-based proteogenomics identifies glioblastoma-specific transposable elements encoding HLA-I-presented peptides. Cell Rep. 39, 110916 (2022).
Zhu, X., Fang, H., Gladysz, K., Barbour, J. A. & Wong, J. W. H. Overexpression of transposable elements is associated with immune evasion and poor outcome in colorectal cancer. Eur. J. Cancer 157, 94–107 (2021).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Stone, M. L. et al. Epigenetic therapy activates type I interferon signaling in murine ovarian cancer to reduce immunosuppression and tumor burden. Proc. Natl Acad. Sci. USA 114, E10981–E10990 (2017).
Roulois, D. et al. DNA-demethylating agents target colorectal cancer cells by inducing viral mimicry by endogenous transcripts. Cell 162, 961–973 (2015).
de Cubas, A. A. et al. DNA hypomethylation promotes transposable element expression and activation of immune signaling in renal cell cancer. JCI Insight 5, e137569 (2020).
Goel, S. et al. CDK4/6 inhibition triggers anti-tumour immunity. Nature 548, 471–475 (2017).
McDonald, J. I. et al. Epigenetic therapies in ovarian cancer alter repetitive element expression in a TP53 -dependent manner. Cancer Res. 81, 5176–5189 (2021).
Topper, M. J. et al. Epigenetic therapy ties MYC depletion to reversing immune evasion and treating lung cancer. Cell 171, 1284–1300.e21 (2017).
Brocks, D. et al. DNMT and HDAC inhibitors induce cryptic transcription start sites encoded in long terminal repeats. Nat. Genet. 49, 1052–1060 (2017).
Goyal, A. et al. DNMT and HDAC inhibition induces immunogenic neoantigens from human endogenous retroviral element-derived transcripts. Nat. Commun. 14, 6731 (2023).
Gomez, S. et al. Inhibiting DNA methylation and RNA editing upregulates immunogenic RNA to transform the tumor microenvironment and prolong survival in ovarian cancer. J. Immunother. Cancer 10, e004974 (2022).
Griffin, G. K. et al. Epigenetic silencing by SETDB1 suppresses tumour intrinsic immunogenicity. Nature 595, 309–314 (2021).
Li, H.-T. et al. RNA mis-splicing drives viral mimicry response after DNMTi therapy in SETD2-mutant kidney cancer. Cell Rep. 42, 112016 (2023).
Sheng, W. et al. LSD1 ablation stimulates anti-tumor immunity and enables checkpoint blockade. Cell 174, 549–563.e19 (2018).
Chiappinelli, K. B., Zahnow, C. A., Ahuja, N. & Baylin, S. B. Combining epigenetic and immunotherapy to combat cancer. Cancer Res. 76, 1683–1689 (2016).
Morel, D., Jeffery, D., Aspeslagh, S., Almouzni, G. & Postel-Vinay, S. Combining epigenetic drugs with other therapies for solid tumours—past lessons and future promise. Nat. Rev. Clin. Oncol. 17, 91–107 (2020).
Licht, J. D. & Bennett, R. L. Leveraging epigenetics to enhance the efficacy of immunotherapy. Clin. Epigenet. 13, 115 (2021).
Belancio, V. P., Roy-Engel, A. M., Pochampally, R. R. & Deininger, P. Somatic expression of LINE-1 elements in human tissues. Nucleic Acids Res. 38, 3909–3922 (2010).
Gardner, E. J. et al. The Mobile Element Locator Tool (MELT): population-scale mobile element discovery and biology. Genome Res. 27, 1916–1929 (2017).
Thung, D. T. et al. Mobster: accurate detection of mobile element insertions in next generation sequencing data. Genome Biol. 15, 488 (2014).
Chu, C. et al. Comprehensive identification of transposable element insertions using multiple sequencing technologies. Nat. Commun. 12, 3836 (2021).
Mohamed, M. et al. TrEMOLO: accurate transposable element allele frequency estimation using long-read sequencing data combining assembly and mapping-based approaches. Genome Biol. 24, 63 (2023).
Disdero, E. & Filée, J. LoRTE: detecting transposon-induced genomic variants using low coverage PacBio long read sequences. Mob. DNA 8, 5 (2017).
Fujimoto, A. et al. Whole-genome sequencing with long reads reveals complex structure and origin of structural variation in human genetic variations and somatic mutations in cancer. Genome Med. 13, 65 (2021).
Jin, Y., Tam, O. H., Paniagua, E. & Hammell, M. TEtranscripts: a package for including transposable elements in differential expression analysis of RNA-seq datasets. Bioinformatics 31, 3593–3599 (2015).
Jeong, H.-H., Yalamanchili, H. K., Guo, C., Shulman, J. M. & Liu, Z. An ultra-fast and scalable quantification pipeline for transposable elements from next generation sequencing data. Pacif. Symp. Biocomput. 23, 168–179 (2018).
Lerat, E., Fablet, M., Modolo, L., Lopez-Maestre, H. & Vieira, C. TEtools facilitates big data expression analysis of transposable elements and reveals an antagonism between their activity and that of piRNA genes. Nucleic Acids Res. 45, e17 (2017).
Criscione, S. W., Zhang, Y., Thompson, W., Sedivy, J. M. & Neretti, N. Transcriptional landscape of repetitive elements in normal and cancer human cells. BMC Genom. 15, 583 (2014).
He, J. et al. Identifying transposable element expression dynamics and heterogeneity during development at the single-cell level with a processing pipeline scTE. Nat. Commun. 12, 1456 (2021).
Yang, W. R., Ardeljan, D., Pacyna, C. N., Payer, L. M. & Burns, K. H. SQuIRE reveals locus-specific regulation of interspersed repeat expression. Nucleic Acids Res. 47, e27 (2019).
Shao, W. & Wang, T. Transcript assembly improves expression quantification of transposable elements in single-cell RNA-seq data. Genome Res. 31, 88–100 (2021).
Bendall, M. L. et al. Telescope: characterization of the retrotranscriptome by accurate estimation of transposable element expression. PLOS Comput. Biol. 15, e1006453 (2019).
McKerrow, W. & Fenyö, D. L1EM: a tool for accurate locus specific LINE-1 RNA quantification. Bioinformatics 36, 1167–1173 (2020).
Maeng, J. H., Jang, H. J., Du, A. Y., Tzeng, S.-C. & Wang, T. Using long-read CAGE sequencing to profile cryptic-promoter-derived transcripts and their contribution to the immunopeptidome. Genome Res. 33, 2143–2155 (2023).
Berrens, R. V. et al. Locus-specific expression of transposable elements in single cells with CELLO-seq. Nat. Biotechnol. 40, 546–554 (2022).
Bourque, G. et al. Ten things you should know about transposable elements. Genome Biol. 19, 199 (2018).
Gualandi, N. et al. Meta-analysis suggests that intron retention can affect quantification of transposable elements from RNA-seq data. Biology 11, 826 (2022).
Faulkner, G. J. Elevated L1 expression in ataxia telangiectasia likely explained by an RNA-seq batch effect. Neuron 111, 610–611 (2023).
Ansaloni, F., Gualandi, N., Esposito, M., Gustincich, S. & Sanges, R. TEspeX: consensus-specific quantification of transposable element expression preventing biases from exonized fragments. Bioinformatics 38, 4430–4433 (2022).
Smit, A. F. A., Hubley, R. & Green, P. RepeatMasker Open-4.0 (2013–2015). ISB http://www.repeatmasker.org (2015).
Babaian, A. et al. LIONS: analysis suite for detecting and quantifying transposable element initiated transcription from RNA-seq. Bioinformatics 35, 3839–3841 (2019).
Lanciano, S. & Cristofari, G. Measuring and interpreting transposable element expression. Nat. Rev. Genet. 21, 721–736 (2020).
Pinson, M.-E., Pogorelcnik, R., Court, F., Arnaud, P. & Vaurs-Barrière, C. CLIFinder: identification of LINE-1 chimeric transcripts in RNA-seq data. Bioinformatics 34, 688–690 (2018).
Sexton, C. E. & Han, M. V. Paired-end mappability of transposable elements in the human genome. Mob. DNA 10, 29 (2019).
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Kershaw, M. H. et al. Immunization against endogenous retroviral tumor-associated antigens. Cancer Res. 61, 7920–7924 (2001).
Wang-Johanning, F. et al. Human endogenous retrovirus K triggers an antigen-specific immune response in breast cancer patients. Cancer Res. 68, 5869–5877 (2008).
Mullins, C. S. & Linnebacher, M. Endogenous retrovirus sequences as a novel class of tumor-specific antigens: an example of HERV-H env encoding strong CTL epitopes. Cancer Immunol. Immunother. 61, 1093–1100 (2012).
Agrawal, A., Eastman, Q. M. & Schatz, D. G. Transposition mediated by RAG1 and RAG2 and its implications for the evolution of the immune system. Nature 394, 744–751 (1998).
Lavialle, C. et al. Paleovirology of ‘syncytins’, retroviral env genes exapted for a role in placentation. Phil. Trans. R. Soc. B 368, 20120507 (2013).
Ferreira, L. M. R. et al. A distant trophoblast-specific enhancer controls HLA-G expression at the maternal–fetal interface. Proc. Natl Acad. Sci. USA 113, 5364–5369 (2016).
Emera, D. & Wagner, G. P. Transformation of a transposon into a derived prolactin promoter with function during human pregnancy. Proc. Natl Acad. Sci. USA 109, 11246–11251 (2012).
Zhang, Y. et al. Transcriptionally active HERV-H retrotransposons demarcate topologically associating domains in human pluripotent stem cells. Nat. Genet. 51, 1380–1388 (2019).
Schmidt, D. et al. Waves of retrotransposon expansion remodel genome organization and CTCF binding in multiple mammalian lineages. Cell 148, 335–348 (2012).
Hermant, C. & Torres-Padilla, M.-E. TFs for TEs: the transcription factor repertoire of mammalian transposable elements. Genes. Dev. 35, 22–39 (2021).
Bourque, G. et al. Evolution of the mammalian transcription factor binding repertoire via transposable elements. Genome Res. 18, 1752–1762 (2008).
Sundaram, V. et al. Widespread contribution of transposable elements to the innovation of gene regulatory networks. Genome Res. 24, 1963–1976 (2014).
Wang, T. et al. Species-specific endogenous retroviruses shape the transcriptional network of the human tumor suppressor protein p53. Proc. Natl Acad. Sci. USA 104, 18613–18618 (2007).
Sundaram, V. & Wang, T. Transposable element mediated innovation in gene regulatory landscapes of cells: re-visiting the “gene-battery” model. BioEssays 40, 1700155 (2018).
Rebollo, R., Romanish, M. T. & Mager, D. L. Transposable elements: an abundant and natural source of regulatory sequences for host genes. Annu. Rev. Genet. 46, 21–42 (2012).
Feschotte, C. Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 9, 397–405 (2008).
Chuong, E. B., Elde, N. C. & Feschotte, C. Regulatory activities of transposable elements: from conflicts to benefits. Nat. Rev. Genet. 18, 71–86 (2017).
Yue, F. et al. A comparative encyclopedia of DNA elements in the mouse genome. Nature 515, 355–364 (2014).
Choudhary, M. N. K., Quaid, K., Xing, X., Schmidt, H. & Wang, T. Widespread contribution of transposable elements to the rewiring of mammalian 3D genomes. Nat. Commun. 14, 634 (2023).
Lowdon, R. F., Jang, H. S. & Wang, T. Evolution of epigenetic regulation in vertebrate genomes. Trends Genet. 32, 269–283 (2016).
Grillo, G. & Lupien, M. Cancer-associated chromatin variants uncover the oncogenic role of transposable elements. Curr. Opin. Genet. Dev. 74, 101911 (2022).
Acknowledgements
The authors thank members of the Wang laboratory for helpful discussions related to the project. Work performed in the Wang laboratory is supported by NIH grants R01HG007175, U24ES026699, U01HG009391 and UM1DA058219.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to all aspects of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Cancer thanks Geoffrey Faulkner and the other, anonymous, reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Alternative splicing
-
A cellular process whereby exons are joined differently to create alternative isoforms of the same gene that have different but related mRNA transcripts.
- Antibody–drug conjugates
-
A rapidly developing class of therapeutic agents that combine monoclonal antibodies with drugs that can achieve cell-type-specific drug delivery.
- A-to-I editing
-
A cellular process that is catalysed by adenosine deaminases acting on RNA (ADARs), which substitute an adenosine for an inosine on the mRNA molecule. Editing of protein-coding RNAs results in dramatic alterations to protein functions. Deficiencies in A-to-I editing lead to genetic diseases. However, the impact of A-to-I editing on Alu-element RNA sequences is still underexplored.
- Bidirectional transcription
-
Transcription events that occur when there is one promoter on each side of a stretch of DNA sequence, initiating transcription over the sequence between the two promoters.
- Cis-regulatory elements
-
DNA sequences that are binding sites for transcription factors and which modulate gene expression.
- Condensate
-
Micrometre-scale subdomains within cells that concentrate biomolecules (such as transcription factors for specialized functions) but that are not bounded by membranes.
- De novo transcript assembly
-
A computational process that utilizes transcriptomic data to construct transcript exon–intron structures.
- DNase-hypersensitive sites
-
Chromatin regions that are characterized by elevated DNase I cleavage because of more accessible, local spatial distribution of nucleosomes.
- Enhancers
-
Elements in the genome that enable the binding of activators and mediators that can subsequently activate expression of distal genes.
- Exapted
-
An event in which a sequence evolves to a function that is different from the original function.
- Insertional mutagenesis
-
Genetic mutations created by insertions of DNA segments, such as disruption of coding sequences of tumour-suppressor genes by LINE-1 retrotranspositions, that contribute to disease progression such as tumorigenesis.
- Intergenic regions
-
DNA sequences located between genes that do not have coding potential. Parts of intergenic regions may contain functional regulatory elements.
- Long non-coding RNA
-
RNA transcript more than 200 nucleotides in length that does not have translation potential. Long non-coding RNAs have a broad function in remodelling of chromatin structure, RNA splicing and stability, and in regulating protein functions by direct binding.
- Long-read sequencing
-
Third-generation sequencing technology that can generate reads of thousands to hundreds of thousands of bases.
- MHC tetramer
-
An important tool that consists of four major histocompatibility complex (MHC) molecules conjugated to a fluorochrome-labelled streptavidin to evaluate the stability of interactions between MHC molecules, antigens and T cell receptors. This tool helps to detect T cells that can recognize specific antigens.
- Nanopore sequencing
-
One of the long-read sequencing platforms that obtains sequence information at single-molecule resolution by measuring the chemical kinetics of each base when passing each ultralong DNA molecule through a nanopore.
- Neoantigens
-
Tumour-specific antigens presented on MHC complexes to trigger an immune response.
- Promoters
-
A segment of DNA sequence that can be bound by RNA polymerase and other transcription machinery to initiate the process of gene expression.
- Sense–antisense pairing
-
Complementary pairing of two nucleotide molecules.
- Short-read sequencing
-
Also known as next-generation sequencing, short-read sequencing generates reads of hundreds of bases.
- Splice acceptor sites
-
DNA sequences at the 3′ end of introns that terminate the intron and facilitate appropriate splicing.
- Splice donor sites
-
DNA sequences at the 5′ end of introns that mark the beginning of introns and facilitate appropriate splicing.
- Transcription-factor-binding sites
-
A stretch of DNA sequence that transcription factors bind to. Each transcription factor will have its own specific sequence pattern (motif) that it recognizes and binds to.
- Trans-effects
-
Gene expression activity regulated by RNAs and proteins by binding to DNA sequences directly or indirectly.
- V(D)J somatic recombination
-
A cellular process that occurs during the development of T cells and B cells that results in a highly diverse repository of T cell receptors and antibodies, respectively.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liang, Y., Qu, X., Shah, N.M. et al. Towards targeting transposable elements for cancer therapy. Nat Rev Cancer 24, 123–140 (2024). https://doi.org/10.1038/s41568-023-00653-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-023-00653-8