Abstract
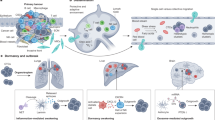
Genetic intratumoural heterogeneity is a natural consequence of imperfect DNA replication. Any two randomly selected cells, whether normal or cancerous, are therefore genetically different. Here, we review the different forms of genetic heterogeneity in cancer and re-analyse the extent of genetic heterogeneity within seven types of untreated epithelial cancers, with particular regard to its clinical relevance. We find that the homogeneity of predicted functional mutations in driver genes is the rule rather than the exception. In primary tumours with multiple samples, 97% of driver-gene mutations in 38 patients were homogeneous. Moreover, among metastases from the same primary tumour, 100% of the driver mutations in 17 patients were homogeneous. With a single biopsy of a primary tumour in 14 patients, the likelihood of missing a functional driver-gene mutation that was present in all metastases was 2.6%. Furthermore, all functional driver-gene mutations detected in these 14 primary tumours were present among all their metastases. Finally, we found that individual metastatic lesions responded concordantly to targeted therapies in 91% of 44 patients. These analyses indicate that the cells within the primary tumours that gave rise to metastases are genetically homogeneous with respect to functional driver-gene mutations, and we suggest that future efforts to develop combination therapies have the potential to be curative.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Heppner, G. H. Tumor heterogeneity. Cancer Res. 44, 2259–2265 (1984).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Martincorena, I. & Campbell, P. J. Somatic mutation in cancer and normal cells. Science 349, 1483–1489 (2015).
Rosenthal, R., McGranahan, N., Herrero, J. & Swanton, C. Deciphering genetic intratumor heterogeneity and its impact on cancer evolution. Annu. Rev. Cancer Biol. 1, 223–240 (2017).
Jones, S. et al. Comparative lesion sequencing provides insights into tumor evolution. Proc. Natl Acad. Sci. USA 105, 4283–4288 (2008).
Engelman, J. A. et al. MET amplification leads to gefitinib resistance in lung cancer by activating ERBB3 signaling. Science 316, 1039–1043 (2007).
Diaz, L. A. Jr et al. The molecular evolution of acquired resistance to targeted EGFR blockade in colorectal cancers. Nature 486, 537–540 (2012).
Flaherty, K. T. et al. Combined BRAF and MEK inhibition in melanoma with BRAF V600 mutations. N. Engl. J. Med. 367, 1694–1703 (2012).
Bozic, I. et al. Evolutionary dynamics of cancer in response to targeted combination therapy. eLife 2, e00747 (2013).
Bozic, I. & Nowak, M. A. Resisting resistance. Annu. Rev. Cancer Biol. 1, 203–221 (2017).
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci. Transl. Med. 7, 283ra54 (2015).
Sottoriva, A., Barnes, C. P. & Graham, T. A. Catch my drift? Making sense of genomic intra-tumour heterogeneity. Biochim. Biophys. Acta Rev. Cancer 1867, 95–100 (2017).
Tamborero, D. et al. Cancer Genome Interpreter annotates the biological and clinical relevance of tumor alterations. Genome Med. 10, 25 (2018).
Reiter, J. G. et al. Minimal functional driver gene heterogeneity among untreated metastases. Science 361, 1033–1037 (2018).
Tokheim, C. & Karchin, R. Enhanced context reveals the scope of somatic missense mutations driving human cancers. Cell Syst. 9, 1–15 (2019).
Vogelstein, B. & Kinzler, K. W. The path to cancer-three strikes and you’re out. N. Engl. J. Med. 373, 1895–1898 (2015).
Tomasetti, C., Marchionni, L., Nowak, M. A., Parmigiani, G. & Vogelstein, B. Only three driver gene mutations are required for the development of lung and colorectal cancers. Proc. Natl Acad. Sci. USA 112, 118–123 (2015).
Hruban, R. H., Goggins, M., Parsons, J. & Kern, S. E. Progression model for pancreatic cancer. Clin. Cancer Res. 6, 2969–2972 (2000).
Cross, W. et al. The evolutionary landscape of colorectal tumorigenesis. Nat. Ecol. Evol. 2, 1661–1672 (2018).
Makohon-Moore, A. P. et al. Precancerous neoplastic cells can move through the pancreatic ductal system. Nature 561, 201–205 (2018).
Saito, T. et al. A temporal shift of the evolutionary principle shaping intratumor heterogeneity in colorectal cancer. Nat. Commun. 9, 2884 (2018).
Bozic, I. et al. Accumulation of driver and passenger mutations during tumor progression. Proc. Natl Acad. Sci. USA 107, 18545–18550 (2010).
Bozic, I., Gerold, J. M. & Nowak, M. A. Quantifying clonal and subclonal passenger mutations in cancer evolution. PLoS Comput. Biol. 12, e1004731 (2016).
Sun, R. et al. Between-region genetic divergence reflects the mode and tempo of tumor evolution. Nat. Genet. 49, 1015–1024 (2017).
Williams, M. J. et al. Quantification of subclonal selection in cancer from bulk sequencing data. Nat. Genet. 50, 895–903 (2018).
Wodarz, D. & Komarova, N. L. Dynamics of Cancer: Mathematical Foundations of Oncology (World Scientific Publishing, 2014).
Altrock, P. M., Liu, L. L. & Michor, F. The mathematics of cancer: integrating quantitative models. Nat. Rev. Cancer 15, 730–745 (2015).
Amikura, K., Kobari, M. & Matsuno, S. The time of occurrence of liver metastasis in carcinoma of the pancreas. Int. J. Pancreatol. 17, 139–146 (1995).
Yachida, S. et al. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature 467, 1114–1117 (2010).
Vermeulen, L. et al. Defining stem cell dynamics in models of intestinal tumor initiation. Science 342, 995–998 (2013).
Cannataro, V. L., Gaffney, S. G. & Townsend, J. P. Effect sizes of somatic mutations in cancer. J. Natl Cancer Inst. 110, 1171–1177 (2018).
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Reiter, J. G., Bozic, I., Allen, B., Chatterjee, K. & Nowak, M. A. The effect of one additional driver mutation on tumor progression. Evol. Appl. 6, 34–45 (2013).
Reiter, J. G., Bozic, I., Chatterjee, K. & Nowak, M. A. TTP: Tool for Tumor Progression in Computer Aided Verification: 25th International Conference, CAV 2013 (eds Sharygina, N. & Veith, H.) 101–106 (Springer, 2013).
Bailey, M. H. et al. Comprehensive characterization of cancer driver genes and mutations. Cell 173, 371–385.e18 (2018).
Sanchez-Vega, F. et al. Oncogenic signaling pathways in the cancer genome atlas. Cell 173, 321–337 (2018).
Landau, D. A. et al. Mutations driving CLL and their evolution in progression and relapse. Nature 526, 525–530 (2015).
Priestley, P. et al. Pan-cancer whole genome analyses of metastatic solid tumors. Preprint at bioRxiv https://doi.org/10.1101/415133 (2019).
Gerstung, M. et al. The evolutionary history of 2,658 cancers. Preprint at bioRxiv https://doi.org/10.1101/161562 (2018).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Kurman, R. J. & Shih, I.-M. The origin and pathogenesis of epithelial ovarian cancer—a proposed unifying theory. Am. J. Surg. Pathol. 34, 433–443 (2010).
Murphy, S. J. et al. Genetic alterations associated with progression from pancreatic intraepithelial neoplasia to invasive pancreatic tumor. Gastroenterology 145, 1098–1109 (2013).
Reiter, J. G. & Iacobuzio-Donahue, C. A. Pancreatic carcinogenesis—several small steps or one giant leap? Nat. Rev. Gastroenterol. Hepatol. 14, 7–8 (2017).
Teixeira, V. H. et al. Deciphering the genomic, epigenomic, and transcriptomic landscapes of pre-invasive lung cancer lesions. Nat. Med. 25, 517–525 (2019).
Lengauer, C., Kinzler, K. W. & Vogelstein, B. Genetic instabilities in human cancers. Nature 396, 643–649 (1998).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Bielski, C. M. et al. Genome doubling shapes the evolution and prognosis of advanced cancers. Nat. Genet. 50, 1189–1195 (2018).
Bolhaqueiro, A. C. F. et al. Ongoing chromosomal instability and karyotype evolution in human colorectal cancer organoids. Nat. Genet. 51, 824–834 (2019).
Massagué, J. & Obenauf, A. C. Metastatic colonization by circulating tumour cells. Nature 529, 298–306 (2016).
Jung, S.-H. et al. Whole-exome sequencing identifies recurrent AKT1 mutations in sclerosing hemangioma of lung. Proc. Natl Acad. Sci. USA 113, 10672–10677 (2016).
Curtius, K., Wright, N. A. & Graham, T. A. Evolution of premalignant disease. Cold Spring Harb. Perspect. Med. 7, a026542 (2017).
Kuboki, Y. et al. Single-cell sequencing defines genetic heterogeneity in pancreatic cancer precursor lesions. J. Pathol. 247, 347–356 (2019).
Winters, I. P., Murray, C. W. & Winslow, M. M. Towards quantitative and multiplexed in vivo functional cancer genomics. Nat. Rev. Genet. 19, 741–755 (2018).
Jamal-Hanjani, M. et al. Tracking the evolution of non-small-cell lung cancer. N. Engl. J. Med. 376, 2109–2121 (2017).
Sondka, Z. et al. The COSMIC cancer gene census: describing genetic dysfunction across all human cancers. Nat. Rev. Cancer 18, 696–705 (2018).
Blokzijl, F. et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature 538, 260–264 (2016).
Bae, T. et al. Different mutational rates and mechanisms in human cells at pregastrulation and neurogenesis. Science 359, 550–555 (2018).
Lodato, M. A. et al. Aging and neurodegeneration are associated with increased mutations in single human neurons. Science 359, 555–559 (2018).
Martincorena, I. et al. Somatic mutant clones colonize the human esophagus with age. Science 362, 911–917 (2018).
Yokoyama, A. et al. Age-related remodelling of oesophageal epithelia by mutated cancer drivers. Nature 565, 312–317 (2019).
Maley, C. C. et al. Classifying the evolutionary and ecological features of neoplasms. Nat. Rev. Cancer 17, 605–619 (2017).
Durrett, R., Foo, J., Leder, K., Mayberry, J. & Michor, F. Intratumor heterogeneity in evolutionary models of tumor progression. Genetics 188, 461–477 (2011).
Almendro, V. et al. Inference of tumor evolution during chemotherapy by computational modeling and in situ analysis of genetic and phenotypic cellular diversity. Cell Rep. 6, 514–527 (2014).
Makohon-Moore, A. P. et al. Limited heterogeneity of known driver gene mutations among the metastases of individual pancreatic cancer patients. Nat. Genet. 49, 358–366 (2017).
Rosenbloom, D. I. S., Camara, P. G., Chu, T. & Rabadan, R. Evolutionary scalpels for dissecting tumor ecosystems. Biochim. Biophys. Acta 1867, 69–83 (2017).
Maley, C. C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat. Genet. 38, 468–473 (2006).
Lemery, S., Keegan, P. & Pazdur, R. First FDA approval agnostic of cancer site-when a biomarker defines the indication. N. Engl. J. Med. 377, 1409–1412 (2017).
Drilon, A. et al. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N. Engl. J. Med. 378, 731–739 (2018).
Makohon-Moore, A. & Iacobuzio-Donahue, C. A. Pancreatic cancer biology and genetics from an evolutionary perspective. Nat. Rev. Cancer 16, 553–565 (2016).
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2018. CA Cancer J. Clin. 68, 7–30 (2018).
Andor, N. et al. Pan-cancer analysis of the extent and consequences of intra-tumor heterogeneity. Nat. Med. 22, 105–113 (2016).
Lacroix, M. et al. A multivariate analysis of 416 patients with glioblastoma multiforme: prognosis, extent of resection, and survival. J. Neurosurg. 95, 190–198 (2001).
Francis, J. M. et al. EGFR variant heterogeneity in glioblastoma resolved through single-nucleus sequencing. Cancer Discov. 4, 956–971 (2014).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Bashashati, A. et al. Distinct evolutionary trajectories of primary high-grade serous ovarian cancers revealed through spatial mutational profiling. J. Pathol. 231, 21–34 (2013).
Gerlinger, M. et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat. Genet. 46, 225–233 (2014).
de Bruin, E. C. et al. Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346, 251–256 (2014).
Yates, L. R. et al. Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat. Med. 21, 751–759 (2015).
Kim, T.-M. et al. Subclonal genomic architectures of primary and metastatic colorectal cancer based on intratumoral genetic heterogeneity. Clin. Cancer Res. 21, 4461–4472 (2015).
McPherson, A. et al. Divergent modes of clonal spread and intraperitoneal mixing in high-grade serous ovarian cancer. Nat. Genet. 48, 758–767 (2016).
Zhang, J. et al. Intratumor heterogeneity in localized lung adenocarcinomas delineated by multiregion sequencing. Science 346, 256–259 (2014).
Reiter, J. G. et al. Reconstructing metastatic seeding patterns of human cancers. Nat. Commun. 8, 14114 (2017).
Shi, W. et al. Reliability of whole-exome sequencing for assessing intratumor genetic heterogeneity. Cell Rep. 25, 1446–1457 (2018).
Zare, F., Dow, M., Monteleone, N., Hosny, A. & Nabavi, S. An evaluation of copy number variation detection tools for cancer using whole exome sequencing data. BMC Bioinformatics 18, 286 (2017).
Salari, R. et al. Inference of tumor phylogenies with improved somatic mutation discovery. J. Comput. Biol. 20, 933–944 (2013).
Zhang, A. W. et al. Interfaces of malignant and immunologic clonal dynamics in ovarian cancer. Cell 173, 1755–1769 (2018).
Ryser, M. D., Min, B.-H., Siegmund, K. D. & Shibata, D. Spatial mutation patterns as markers of early colorectal tumor cell mobility. Proc. Natl Acad. Sci. USA 115, 5774–5779 (2018).
Pectasides, E. et al. Genomic heterogeneity as a barrier to precision medicine in gastroesophageal adenocarcinoma. Cancer Discov. 8, 37–48 (2018).
LiFD (Likely Functional Driver). https://github.com/johannesreiter/LiFD (2019).
Chakravarty, D. et al. OncoKB: a precision oncology knowledge base. JCO Precis. Oncol. 1, 1–16 (2017).
Chang, M. T. et al. Identifying recurrent mutations in cancer reveals widespread lineage diversity and mutational specificity. Nat. Biotechnol. 34, 155–163 (2016).
Forbes, S. et al. COSMIC: high-resolution cancer genetics using the catalogue of somatic mutations in cancer. Curr. Protoc. Hum. Genet. 91, 1–37 (2016).
Masica, D. L. et al. CRAVAT 4: cancer-related analysis of variants toolkit. Cancer Res. 77, e35–e38 (2017).
Shihab, H. A., Gough, J., Cooper, D. N., Day, I. N. M. & Gaunt, T. R. Predicting the functional consequences of cancer-associated amino acid substitutions. Bioinformatics 29, 1504–1510 (2013).
Mao, Y. et al. CanDrA: cancer-specific driver missense mutation annotation with optimized features. PLoS ONE 8, e77945 (2013).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Løes, I. M. et al. Impact of KRAS, BRAF, PIK3CA, TP53 status and intraindividual mutation heterogeneity on outcome after liver resection for colorectal cancer metastases. Int. J. Cancer 139, 647–656 (2016).
Smith, J. C. & Sheltzer, J. M. Systematic identification of mutations and copy number alterations associated with cancer patient prognosis. eLife 7, e39217 (2018).
Schwarz, R. F. et al. Spatial and temporal heterogeneity in high-grade serous ovarian cancer: a phylogenetic analysis. PLoS Med. 12, e1001789 (2015).
Gokulan, R. C., Garcia-Buitrago, M. T. & Zaika, A. I. From genetics to signaling pathways: molecular pathogenesis of esophageal adenocarcinoma. Biochim. Biophys. Acta Rev. Cancer 1872, 37–48 (2019).
Naxerova, K. & Jain, R. K. Using tumour phylogenetics to identify the roots of metastasis in humans. Nat. Rev. Clin. Oncol. 12, 258–272 (2015).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur. J. Cancer 45, 228–247 (2009).
Gibson, W. J. et al. The genomic landscape and evolution of endometrial carcinoma progression and abdominopelvic metastasis. Nat. Genet. 48, 848–855 (2016).
Brown, D. et al. Phylogenetic analysis of metastatic progression in breast cancer using somatic mutations and copy number aberrations. Nat. Commun. 8, 14944 (2017).
Hong, M. K. H. et al. Tracking the origins and drivers of subclonal metastatic expansion in prostate cancer. Nat. Commun. 6, 6605 (2015).
Tokheim, C. J., Papadopoulos, N., Kinzler, K. W., Vogelstein, B. & Karchin, R. Evaluating the evaluation of cancer driver genes. Proc. Natl Acad. Sci. USA 113, 14330–14335 (2016).
Turajlic, S. & Swanton, C. Metastasis as an evolutionary process. Science 352, 169–175 (2016).
Kumar, A. et al. Substantial interindividual and limited intraindividual genomic diversity among tumors from men with metastatic prostate cancer. Nat. Med. 22, 369–378 (2016).
Gundem, G. et al. The evolutionary history of lethal metastatic prostate cancer. Nature 520, 353–357 (2015).
Zhao, Z.-M. et al. Early and multiple origins of metastatic lineages within primary tumors. Proc. Natl Acad. Sci. USA 113, 2140–2145 (2016).
Hunter, K. W., Amin, R., Deasy, S., Ha, N.-H. & Wakefield, L. Genetic insights into the morass of metastatic heterogeneity. Nat. Rev. Cancer 18, 211–223 (2018).
Sharma, S. V., Bell, D. W., Settleman, J. & Haber, D. A. Epidermal growth factor receptor mutations in lung cancer. Nat. Rev. Cancer 7, 169–181 (2007).
Juric, D. et al. Convergent loss of PTEN leads to clinical resistance to a PI(3)Kα inhibitor. Nature 518, 240–244 (2015).
Khan, K. H. et al. Longitudinal liquid biopsy and mathematical modeling of clonal evolution forecast time to treatment failure in the PROSPECT-C phase II colorectal cancer clinical trial. Cancer Discov. 8, 1270–1285 (2018).
Kobayashi, S. et al. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 352, 786–792 (2005).
Goyal, L. et al. Polyclonal secondary FGFR2 mutations drive acquired resistance to FGFR inhibition in patients with FGFR2 fusion-positive cholangiocarcinoma. Cancer Discov. 7, 252–263 (2017).
Vakiani, E. et al. Comparative genomic analysis of primary versus metastatic colorectal carcinomas. J. Clin. Oncol. 30, 2956–2962 (2012).
Brannon, A. R. et al. Comparative sequencing analysis reveals high genomic concordance between matched primary and metastatic colorectal cancer lesions. Genome Biol. 15, 454 (2014).
Yatabe, Y., Matsuo, K. & Mitsudomi, T. Heterogeneous distribution of EGFR mutations is extremely rare in lung adenocarcinoma. J. Clin. Oncol. 29, 2972–2977 (2011).
Colombino, M. et al. BRAF/NRAS mutation frequencies among primary tumors and metastases in patients with melanoma. J. Clin. Oncol. 30, 2522–2529 (2012).
Boursault, L. et al. Tumor homogeneity between primary and metastatic sites for BRAF status in metastatic melanoma determined by immunohistochemical and molecular testing. PLoS ONE 8, e70826 (2013).
Ribas, A. et al. Combination of vemurafenib and cobimetinib in patients with advanced BRAFV600-mutated melanoma: a phase 1b study. Lancet Oncol. 15, 954–965 (2014).
Algazi, A. P. et al. SWOG S1221: a phase 1 dose escalation study co-targeting MAPK-dependent and MAPK-independent BRAF inhibitor resistance in BRAF mutant advanced solid tumors with dabrafenib, trametinib, and GSK2141795 (ClinicalTrials.gov NCT01902173). J. Clin. Oncol. 35, 2578 (2017).
Kurzrock, R. et al. The VEGF receptor tyrosine kinase inhibitor pazopanib in combination with the MEK inhibitor trametinib in advanced solid tumors and differentiated thyroid cancers. Clin. Cancer Res. https://doi.org/10.1158/1078-0432.CCR-18-1881 (2019).
Olive, K. P. et al. Inhibition of Hedgehog signaling enhances delivery of chemotherapy in a mouse model of pancreatic cancer. Science 324, 1457–1461 (2009).
Jain, R. K. Normalizing tumor microenvironment to treat cancer: bench to bedside to biomarkers. J. Clin. Oncol. 31, 2205–2218 (2013).
Pommier, A. et al. Unresolved endoplasmic reticulum stress engenders immune-resistant, latent pancreatic cancer metastases. Science 360, eaao4908 (2018).
Turajlic, S. et al. Tracking cancer evolution reveals constrained routes to metastases: TRACERx renal. Cell 173, 581–594.e12 (2018).
Naxerova, K. et al. Origins of lymphatic and distant metastases in human colorectal cancer. Science 357, 55–60 (2017).
Schwartz, R. & Schäffer, A. A. The evolution of tumour phylogenetics: principles and practice. Nat. Rev. Genet. 18, 213–229 (2017).
Mitchell, T. J. et al. Timing the landmark events in the evolution of clear cell renal cell cancer: TRACERx renal. Cell 173, 611–623.e17 (2018).
Beerenwinkel, N., Schwarz, R. F., Gerstung, M. & Markowetz, F. Cancer evolution: mathematical models and computational inference. Syst. Biol. 64, e1–e25 (2015).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
Santarius, T., Shipley, J., Brewer, D., Stratton, M. R. & Cooper, C. S. A census of amplified and overexpressed human cancer genes. Nat. Rev. Cancer 10, 59–64 (2010).
Xue, W. et al. A cluster of cooperating tumor-suppressor gene candidates in chromosomal deletions. Proc. Natl Acad. Sci. USA 109, 8212–8217 (2012).
Solimini, N. L. et al. Recurrent hemizygous deletions in cancers may optimize proliferative potential. Science 337, 104–109 (2012).
Knouse, K. A., Davoli, T., Elledge, S. J. & Amon, A. Aneuploidy in cancer: Seq-ing answers to old questions. Annu. Rev. Cancer Biol. 1, 335–354 (2017).
Rheinbay, E. et al. Recurrent and functional regulatory mutations in breast cancer. Nature 547, 55–60 (2017).
Rheinbay, E. et al. Discovery and characterization of coding and non-coding driver mutations in more than 2,500 whole cancer genomes. Preprint at bioRxiv https://doi.org/10.1101/237313 (2017).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
El-Kebir, M., Satas, G. & Raphael, B. J. Inferring parsimonious migration histories for metastatic cancers. Nat. Genet. 50, 718–726 (2018).
Heyde, A., Reiter, J. G., Naxerova, K. & Nowak, M. A. Consecutive seeding and transfer of genetic diversity in metastasis. Proc. Natl Acad. Sci. USA 116, 14129–14137 (2019).
Acknowledgements
The authors thank all authors of the original publications for enabling this re-analysis by sharing their sequencing data — in particular, A. Bass, D. Brown, C. Sotiriou, C. Curtis, W. Gibson, E. Hoivik, M. Cmero, C. Hovens, T.-M. Kim, S.-H. Lee, M. Ryser, S. Shah, D. Shibata, M. Stachler, R. Sun and A. Zhang. This study was supported by the US National Institutes of Health grants R00 CA229991 (J.G.R.), CA179991 (C.A.I.-D.), F31 CA180682 (A.P.M.-M.), T32 CA160001-06 (A.P.M.-M.) and CA43460 (B.V.), as well as by the Lustgarten Foundation for Pancreatic Cancer Research, The Sol Goldman Pancreatic Cancer Research Center, the Virginia and D. K. Ludwig Fund for Cancer Research, an Erwin Schrödinger fellowship (J.G.R.; Austrian Science Fund FWF J-3996), a Landry Cancer Biology fellowship (J.M.G.) and the Office of Naval Research grant N00014-16-1-2914.
Author information
Authors and Affiliations
Contributions
All authors researched data for this article. J.G.R., M.B., J.M.G., A.P.M.-M., C.A.I.-D., N.S.A., K.W.K., M.A.N. and B.V. substantially contributed to discussion of the content. J.G.R., M.B., N.S.A., M.A.N. and B.V. wrote the article. All authors reviewed and/or edited the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
K.W.K. and B.V. are founders of Personal Genome Diagnostics and Thrive, as well as advisers to Sysmex, Eisai, CAGE and Neophore. B.V. is also an adviser to Nexus. These companies and others have licensed technologies related to the work described in this paper from Johns Hopkins University. Some of these licences are associated with equity or royalty payments to K.W.K. and B.V. The terms of these arrangements are being managed by Johns Hopkins University in accordance with its conflict-of-interest policies. The other authors declare no competing interests.
Additional information
Peer review information
Nature Reviews Cancer thanks T. Davoli, M. Gerstung and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Glossary
- Subclonal
-
Mutations present in only a subset of the tumour’s cells. They are sometimes described as ‘branched’ because they occur on a branch of the tree when the evolutionary trajectory of the tumour is assessed.
- Clonal
-
Mutations present in virtually all cells of the tumour. They are also called ‘truncal’ because they are in the trunk of the tumour evolutionary tree.
- Clonal sweep
-
A mechanism through which a subpopulation sweeps through a tumour and drives all other competing subpopulations to extinction.
- Simpson index
-
An index denoting the probability that two randomly chosen cancer cells would belong to the same subclone.
- Shannon index
-
An index describing the uncertainty of predicting the subclone of a randomly chosen cancer cell.
- Jaccard similarity coefficient
-
A measure defined as the ratio of the number of shared mutations to all mutations in two samples.
- Selective bottlenecks
-
The scenario in which a decrease of the tumour size (for example, due to therapy) leads to a decrease in genetic diversity and an increase in the prevalence of some subclones in the tumour.
- Response evaluation criteria in solid tumours
-
(RECIST). A standardized measure of solid tumour response to a therapy.
Rights and permissions
About this article
Cite this article
Reiter, J.G., Baretti, M., Gerold, J.M. et al. An analysis of genetic heterogeneity in untreated cancers. Nat Rev Cancer 19, 639–650 (2019). https://doi.org/10.1038/s41568-019-0185-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41568-019-0185-x
This article is cited by
-
Tumor-derived cell-free DNA and circulating tumor cells: partners or rivals in metastasis formation?
Clinical and Experimental Medicine (2024)
-
ACT-Discover: identifying karyotype heterogeneity in pancreatic cancer evolution using ctDNA
Genome Medicine (2023)
-
Spatial biology of cancer evolution
Nature Reviews Genetics (2023)
-
Unraveling the Drivers of Tumorigenesis in the Context of Evolution: Theoretical Models and Bioinformatics Tools
Journal of Molecular Evolution (2023)
-
Determination of the androgen receptor status of disseminated tumor cells in primary breast cancer patients
Archives of Gynecology and Obstetrics (2023)