Abstract
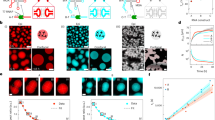
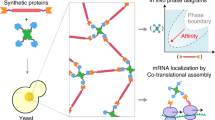
The assembly of protein machines in cells is precise, rapid, and coupled to protein synthesis with regulation in space and time. The assembly of natural and synthetic nanomachines could be similarly controlled by genetic programming outside the cell. Here, we present quasi-two-dimensional (2D) silicon compartments that enable programming of protein assembly lines by local synthesis from surface-immobilized DNA brushes. Using this platform, we studied the autonomous synthesis and assembly of a structural complex from a bacteriophage and a bacterial RNA-synthesizing machine. Local synthesis and surface capture of complexes provided high assembly yield and sensitive detection of spatially resolved assembly intermediates, with the 3D geometry of the compartment and the 2D pattern of brushes dictating the yield and mode of assembly steps. Localized synthesis of proteins in a single gene brush enhances their interactions, and displacement of their genes in separated brushes leads to step-by-step surface assembly. This methodology enables spatial regulation of protein synthesis, and deciphering, reconstruction and design of biological machine assembly lines.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The data necessary to interpret, replicate and build upon this work appears in the article and its Supplementary information. Additional data can be made available from the corresponding authors upon reasonable request.
Code availability
The information needed for the computer simulation in Fig. 2 appears in the article and its Supplementary information.
References
Gu, H., Chao, J., Xiao, S.-J. & Seeman, N. C. A proximity-based programmable DNA nanoscale assembly line. Nature 465, 202–205 (2010).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Bath, J. & Turberfield, A. J. DNA nanomachines. Nat. Nanotechnol. 2, 275–284 (2007).
Leung, K., Chakraborty, K., Saminathan, A. & Krishnan, Y. A DNA nanomachine chemically resolves lysosomes in live cells. Nat. Nanotechnol. 14, 176–183 (2019).
Douglas, S. M., Bachelet, I. & Church, G. M. A logic-gated nanorobot for targeted transport of molecular payloads. Science 335, 831–834 (2012).
Schwarz-Schilling, M. et al. Optimized assembly of a multifunctional RNA-protein nanostructure in a cell-free gene expression system. Nano Lett. 18, 2650–2657 (2018).
Freeman, R. et al. Reversible self-assembly of superstructured networks. Science 362, 808–813 (2018).
Strackharn, M., Pippig, D. A., Meyer, P., Stahl, S. W. & Gaub, H. E. Nanoscale arrangement of proteins by single-molecule cut-and-paste. J. Am. Chem. Soc. 134, 15193–15196 (2012).
Daube, S. S. & Bar-Ziv, R. H. Protein nanomachines assembly modes: cell-free expression and biochip perspectives. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 5, 613–628 (2013).
Shieh, Y.-W. et al. Operon structure and cotranslational subunit association direct protein assembly in bacteria. Science 350, 678–680 (2015).
Holt, C. E., Martin, K. C. & Schuman, E. M. Local translation in neurons: visualization and function. Nat. Struct. Mol. Biol. 26, 557–566 (2019).
Minton, A. P. Implications of macromolecular crowding for protein assembly. Curr. Opin. Struct. Biol. 10, 34–39 (2000).
Daube, S. S., Arad, T. & Bar-Ziv, R. Cell-free co-synthesis of protein nanoassemblies: tubes, rings, and doughnuts. Nano Lett. 7, 638–641 (2007).
Asahara, H. & Chong, S. In vitro genetic reconstruction of bacterial transcription initiation by coupled synthesis and detection of RNA polymerase holoenzyme. Nucleic Acids Res. 38, e141 (2010).
Matthies, D. et al. Cell-free expression and assembly of ATP synthase. J. Mol. Biol. 413, 593–603 (2011).
Rustad, M., Eastlund, A., Jardine, P. & Noireaux, V. Cell-free TXTL synthesis of infectious bacteriophage T4 in a single test tube reaction. Synth. Biol. 3, ysy002 (2018).
Bracha, D., Karzbrun, E., Daube, S. S. & Bar-Ziv, R. H. Emergent properties of dense DNA phases toward artificial biosystems on a surface. Acc. Chem. Res. 47, 1912–1921 (2014).
Daube, S., Bracha, D., Buxboim, A. & Bar-Ziv, R. H. Compartmentalization by directional gene expression. Proc. Natl Acad. Sci. USA 107, 2836–2841 (2010).
Heyman, Y., Buxboim, A., Wolf, S. G., Daube, S. S. & Bar-Ziv, R. H. Cell-free protein synthesis and assembly on a biochip. Nat. Nanotechnol. 7, 374–378 (2012).
Levy, M., Falkovich, R., Daube, S. S. & Bar-Ziv, R. H. Autonomous synthesis and assembly of a ribosomal subunit on a chip. Sci. Adv. 6, eaaz6020 (2020).
Karzbrun, E., Tayar, A. M., Noireaux, V. & Bar-Ziv, R. H. Programmable on-chip DNA compartments as artificial cells. Science 345, 829–832 (2014).
Tayar, A. M., Karzbrun, E., Noireaux, V. & Bar-Ziv, R. H. Synchrony and pattern formation of coupled genetic oscillators on a chip of artificial cells. Proc. Natl. Acad. Sci. USA 114, 11609–11614 (2017).
Efrat, Y., Tayar, A. M., Daube, S. S., Levy, M. & Bar-Ziv, R. H. Electric-field manipulation of a compartmentalized cell-free gene expression reaction. ACS Synth. Biol. 7, 1829–1833 (2018).
Shin, J., Jardine, P. & Noireaux, V. Genome replication, synthesis, and assembly of the bacteriophage T7 in a single cell-free reaction. ACS Synth. Biol. 1, 408–413 (2012).
Garamella, J., Marshall, R., Rustad, M. & Noireaux, V. The all E. coli TX-TL toolbox 2.0: a platform for cell-free synthetic biology. ACS Synth. Biol. 5, 344–355 (2016).
Yap, M. L. et al. Role of bacteriophage T4 baseplate in regulating assembly and infection. Proc. Natl Acad. Sci. USA 113, 2654–2659 (2016).
Yap, M. L. et al. Sequential assembly of the wedge of the baseplate of phage T4 in the presence and absence of gp11 as monitored by analytical ultracentrifugation. Macromol. Biosci. 10, 808–813 (2010).
Bray, D. & Lay, S. Computer-based analysis of the binding steps in protein complex formation. Proc. Natl Acad. Sci. USA 94, 13493–13498 (1997).
Roy, R. D., Rosenmund, C. & Stefan, M. I. Cooperative binding mitigates the high-dose hook effect. BMC Syst. Biol. 11, 74 (2017).
Murugan, A. et al. Undesired usage and the robust self-assembly of heterogeneous structures. Nat. Commun. 6, 6203 (2015).
Luke, K. et al. Microarray analysis of gene expression during bacteriophage T4 infection. Virology 299, 182–191 (2002).
Marshall, R. & Noireaux, V. Quantitative modeling of transcription and translation of an all-E. coli cell-free system. Sci. Rep. 9, 11980 (2019).
Shimizu, Y. et al. Cell-free translation reconstituted with purified components. Nat. Biotechnol. 19, 751–755 (2001).
Mathew, R. & Chatterji, D. The evolving story of the omega subunit of bacterial RNA polymerase. Trends Microbiol. 14, 450–455 (2006).
Hillebrecht, J. R. & Chong, S. A comparative study of protein synthesis in in vitro systems: from the prokaryotic reconstituted to the eukaryotic extract-based. BMC Biotechnol. 8, 58 (2008).
Erijman, A., Dantes, A., Bernheim, R., Shifman, J. M. & Peleg, Y. Transfer-PCR (TPCR): a highway for DNA cloning and protein engineering. J. Struct. Biol. 175, 171–177 (2011).
Kincade, J. M. & deHaseth, P. L. Bacteriophage lambda promoters pL and pR sequence determinants of in vivo activity and of sensitivity to the DNA gyrase inhibitor, coumermycin. Gene 97, 7–12 (1991).
Buxboim, A., Daube, S. S. & Bar-Ziv, R. Ultradense synthetic gene brushes on a chip. Nano Lett. 9, 909–913 (2009).
Caschera, F. & Noireaux, V. Synthesis of 2.3 mg/ml of protein with an all Escherichia coli cell-free transcription–translation system. Biochimie 99, 162–168 (2014).
He, B. et al. Rapid mutagenesis and purification of phage RNA polymerases. Protein Express. Purif. 9, 142–151 (1997).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Buxboim, A. et al. A single-step photolithographic interface for cell-free gene expression and active biochips. Small 3, 500–510 (2007).
Acknowledgements
We acknowledge funding from the Israel Science Foundation (grant no. 1870/15), the United States–Israel Binational Science Foundation (grant no. 2014400), and the Minerva Foundation (grant no. 712274) for the work on the T4 wedges. We thank the Office of Naval Research (award no. N62909-18-1-2094) for funding the work on RNAP assembly. We thank M. Levy for discussions.
Author information
Authors and Affiliations
Contributions
O.V., S.S.D., R.H.B.-Z. and V.N. conceived the cell-free assembly of phages. O.V., Y.D., S.S.D. and R.H.B.-Z. designed the experiments and analysed the data. O.V. and Y.D. performed the experiments. D.G. and V.N. provided the cell-free E. coli extract. O.V., S.F., S.R. and R.L. did the modelling and computational work. O.V., Y.D., S.F., S.R., S.S.D. and R.H.B.-Z. wrote the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Competition between on-chip solution and scaffolded assembly.
a, Scheme, co-expression and interaction of gp10 and gp7 in solution prior to surface binding sequester the scaffolding of gp7 onto gp10 pre-bound to the surface. b, Dose response of gene-10 fraction in a mixed DNA brush with fixed amount of gene-7. Surface gp11 traps were pre-bound with gp10. Surface bound gp10-7 complexes revealed by post-staining with FL-gp8. Detailed gene composition of all experiments appears in Supplementary Table 3. Individual data points and mean values are shown, error bars represent ± s.d. The number of samples for each data point (b) is listed in Supplementary Table 4.
Supplementary information
Supplementary Information
Supplementary information on computational modelling, Supplementary Figs. 1–11 and Tables 1–4.
Rights and permissions
About this article
Cite this article
Vonshak, O., Divon, Y., Förste, S. et al. Programming multi-protein assembly by gene-brush patterns and two-dimensional compartment geometry. Nat. Nanotechnol. 15, 783–791 (2020). https://doi.org/10.1038/s41565-020-0720-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-020-0720-7
This article is cited by
-
A genetic circuit on a single DNA molecule as an autonomous dissipative nanodevice
Nature Communications (2024)
-
PHEIGES: all-cell-free phage synthesis and selection from engineered genomes
Nature Communications (2024)
-
Computational analysis of protein synthesis, diffusion, and binding in compartmental biochips
Microbial Cell Factories (2023)