Abstract
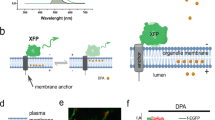
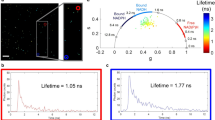
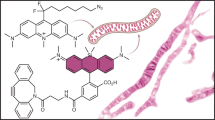
The role of membrane potential in most intracellular organelles remains unexplored because of the lack of suitable tools. Here, we describe Voltair, a fluorescent DNA nanodevice that reports the absolute membrane potential and can be targeted to organelles in live cells. Voltair consists of a voltage-sensitive fluorophore and a reference fluorophore for ratiometry, and acts as an endocytic tracer. Using Voltair, we could measure the membrane potential of different organelles in situ in live cells. Voltair can potentially guide the rational design of biocompatible electronics and enhance our understanding of how membrane potential regulates organelle biology.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The data supporting the plots in this paper and other findings of this study are available from the corresponding author upon reasonable request.
Code availability
The MATLAB code for the voltage clamping experiments are available from the corresponding author upon reasonable request.
References
Alberts, B. et al. Ion Channels and the Electrical Properties of Membranes. in Molecular Biology of the Cell 4th edn (Garland Science, 2002).
Mousavi, S. A. R., Chauvin, A., Pascaud, F., Kellenberger, S. & Farmer, E. E. GLUTAMATE RECEPTOR-LIKE genes mediate leaf-to-leaf wound signalling. Nature 500, 422–426 (2013).
Grabe, M. & Oster, G. Regulation of organelle acidity. J. Gen. Physiol. 117, 329–344 (2001).
Zhong, X. Z. & Dong, X.-P. Lysosome electrophysiology. Methods Cell. Biol. 126, 197–215 (2015).
Meeusen, S., McCaffery, J. M. & Nunnari, J. Mitochondrial fusion intermediates revealed in vitro. Science 305, 1747–1752 (2004).
Ramzan, R., Staniek, K., Kadenbach, B. & Vogt, S. Mitochondrial respiration and membrane potential are regulated by the allosteric ATP-inhibition of cytochrome c oxidase. Biochim. Biophys. Acta 1797, 1672–1680 (2010).
Wang, W. et al. A voltage-dependent K+ channel in the lysosome is required for refilling lysosomal Ca2+ stores. J. Cell Biol. 216, 1715–1730 (2017).
Wang, X. et al. TPC proteins are phosphoinositide-activated sodium-selective ion channels in endosomes and lysosomes. Cell 151, 372–383 (2012).
Dong, X.-P., Wang, X. & Xu, H. TRP channels of intracellular membranes. J. Neurochem. 113, 313–328 (2010).
Srivastava, J., Barber, D. L. & Jacobson, M. P. Intracellular pH sensors: design principles and functional significance. Physiology 22, 30–39 (2007).
Kulkarni, R. U. et al. Voltage-sensitive rhodol with enhanced two-photon brightness. Proc. Natl Acad. Sci. USA 114, 2813–2818 (2017).
Miller, E. W. Small molecule fluorescent voltage indicators for studying membrane potential. Curr. Opin. Chem. Biol. 33, 74–80 (2016).
Dan, K., Veetil, A. T., Chakraborty, K. & Krishnan, Y. DNA nanodevices map enzymatic activity in organelles. Nat. Nanotechnol. 14, 252–259 (2019).
Leung, K., Chakraborty, K., Saminathan, A. & Krishnan, Y. A DNA nanomachine chemically resolves lysosomes in live cells. Nat. Nanotechnol. 14, 176–183 (2019).
Zhao, B. et al. Visualizing intercellular tensile forces by DNA-based membrane molecular probes. J. Am. Chem. Soc. 139, 18182–18185 (2017).
Veetil, A. T. et al. DNA-based fluorescent probes of NOS2 activity in live brains. Proc. Natl Acad. Sci. USA 117, 14694–14702 (2020).
Famulok, M., Hartig, J. S. & Mayer, G. Functional aptamers and aptazymes in biotechnology, diagnostics, and therapy. Chem. Rev. 107, 3715–3743 (2007).
Saha, S., Prakash, V., Halder, S., Chakraborty, K. & Krishnan, Y. A pH-independent DNA nanodevice for quantifying chloride transport in organelles of living cells. Nat. Nanotechnol. 10, 645–651 (2015).
Modi, S., Nizak, C., Surana, S., Halder, S. & Krishnan, Y. Two DNA nanomachines map pH changes along intersecting endocytic pathways inside the same cell. Nat. Nanotechnol. 8, 459–467 (2013).
Rubaiy, H. N. A short guide to electrophysiology and ion channels. J. Pharm. Pharm. Sci. 20, 48–67 (2017).
Lazzari-Dean, J. R., Gest, A. M. & Miller, E. W. Optical estimation of absolute membrane potential using fluorescence lifetime imaging. eLife 8, e44522 (2019).
van Lengerich, B., Rawle, R. J. & Boxer, S. G. Covalent attachment of lipid vesicles to a fluid-supported bilayer allows observation of DNA-mediated vesicle interactions. Langmuir 26, 8666–8672 (2010).
Chakraborty, K., Veetil, A. T., Jaffrey, S. R. & Krishnan, Y. Nucleic acid-based nanodevices in biological imaging. Annu. Rev. Biochem. 85, 349–373 (2016).
Jin, L. et al. Characterization and application of a new optical probe for membrane lipid domains. Biophys. J. 90, 2563–2575 (2006).
Fliegert, R. et al. Modulation of Ca2+ entry and plasma membrane potential by human TRPM4b. FEBS J. 274, 704–713 (2007).
Post, S. R. et al. Class A scavenger receptors mediate cell adhesion via activation of G(i/o) and formation of focal adhesion complexes. J. Lipid Res. 43, 1829–1836 (2002).
Rosenfeld, J. L. et al. Lysosome proteins are redistributed during expression of a GTP-hydrolysis-defective rab5a. J. Cell Sci. 114, 4499–4508 (2001).
Magadán, J. G., Barbieri, M. A., Mesa, R., Stahl, P. D. & Mayorga, L. S. Rab22a regulates the sorting of transferrin to recycling endosomes. Mol. Cell. Biol. 26, 2595–2614 (2006).
Koivusalo, M., Steinberg, B. E., Mason, D. & Grinstein, S. In situ measurement of the electrical potential across the lysosomal membrane using FRET. Traffic 12, 972–982 (2011).
Farsi, Z. et al. Single-vesicle imaging reveals different transport mechanisms between glutamatergic and GABAergic vesicles. Science 351, 981–984 (2016).
Cang, C. et al. mTOR regulates lysosomal ATP-sensitive two-pore Na(+) channels to adapt to metabolic state. Cell 152, 778–790 (2013).
Sun, X. et al. A negative feedback regulation of MTORC1 activity by the lysosomal Ca2+ channel MCOLN1 (mucolipin 1) using a CALM (calmodulin)-dependent mechanism. Autophagy 14, 38–52 (2018).
Hockey, L. N. et al. Dysregulation of lysosomal morphology by pathogenic LRRK2 is corrected by TPC2 inhibition. J. Cell Sci. 128, 232–238 (2015).
Lloyd-Evans, E., Waller-Evans, H., Peterneva, K. & Platt, F. M. Endolysosomal calcium regulation and disease. Biochem. Soc. Trans. 38, 1458–1464 (2010).
Narayanaswamy, N. et al. A pH-correctable, DNA-based fluorescent reporter for organellar calcium. Nat. Methods 16, 95–102 (2019).
Chakraborty, K., Leung, K. & Krishnan, Y. High lumenal chloride in the lysosome is critical for lysosome function. eLife 6, e28862 (2017).
Modi, S. et al. A DNA nanomachine that maps spatial and temporal pH changes inside living cells. Nat. Nanotechnol. 4, 325–330 (2009).
Hover, S. et al. Bunyavirus requirement for endosomal K+ reveals new roles of cellular ion channels during infection. PLoS Pathog. 14, e1006845 (2018).
Cain, C. C., Sipe, D. M. & Murphy, R. F. Regulation of endocytic pH by the Na+,K+-ATPase in living cells. Proc. Natl Acad. Sci. USA 86, 544–548 (1989).
Schapiro, F. B. & Grinstein, S. Determinants of the pH of the Golgi complex. J. Biol. Chem. 275, 21025–21032 (2000).
Gagescu, R. et al. The recycling endosome of Madin-Darby canine kidney cells is a mildly acidic compartment rich in raft components. Mol. Biol. Cell. 11, 2775–2791 (2000).
Numata, M. & Orlowski, J. Molecular cloning and characterization of a novel (Na+,K+)/H+ exchanger localized to the trans-Golgi network. J. Biol. Chem. 276, 17387–17394 (2001).
Raffaello, A., Mammucari, C., Gherardi, G. & Rizzuto, R. Calcium at the center of cell signaling: interplay between endoplasmic reticulum, mitochondria, and lysosomes. Trends Biochem. Sci. 41, 1035–1049 (2016).
Atakpa, P., Thillaiappan, N. B., Mataragka, S., Prole, D. L. & Taylor, C. W. IP3 Receptors preferentially associate with ER-lysosome contact sites and selectively deliver Ca2+ to lysosomes. Cell Rep. 25, 3180–3193.e7 (2018).
Russa, A. D. et al. Microtubule remodeling mediates the inhibition of store-operated calcium entry (SOCE) during mitosis in COS-7 cells. Arch. Histol. Cytol. 71, 249–263 (2008).
Lloyd-Evans, E. On the move, lysosomal CAX drives Ca2+ transport and motility. J. Cell Biol. 212, 755–757 (2016).
Yang, H. H. & St-Pierre, F. Genetically encoded voltage indicators: opportunities and challenges. J. Neurosci. 36, 9977–9989 (2016).
Vaithianathan, T., Henry, D., Akmentin, W. & Matthews, G. Nanoscale dynamics of synaptic vesicle trafficking and fusion at the presynaptic active zone. eLife 5, e13245 (2016).
Jani, M. S., Zou, J., Veetil, A. T. & Krishnan, Y. A DNA-based fluorescent probe maps NOS3 activity with subcellular spatial resolution. Nat. Chem. Biol. 16, 660–666 (2020).
Acarón Ledesma, H. et al. An atlas of nano-enabled neural interfaces. Nat. Nanotechnol. 14, 645–657 (2019).
Prakash, V. et al. Quantitative mapping of endosomal DNA processing by single molecule counting. Angew. Chem. Int. Ed. Engl. 58, 3073–3076 (2019).
Lahann, J. Click Chemistry for Biotechnology and Materials Science (John Wiley & Sons, 2009).
Thekkan, S. et al. A DNA-based fluorescent reporter maps HOCl production in the maturing phagosome. Nat. Chem. Biol. 15, 1165–1172 (2019).
Moore, D. & Dowhan, D. Purification and concentration of DNA from aqueous solutions. Curr. Protoc. Mol. Biol. https://doi.org/10.1002/0471142727.mb0201as59 (2002).
Vonderheit, A. & Helenius, A. Rab7 associates with early endosomes to mediate sorting and transport of Semliki forest virus to late endosomes. PLoS Biol. 3, e233 (2005).
Grimm, C., Vierock, J., Hegemann, P. & Wietek, J. Whole-cell patch-clamp recordings for electrophysiological determination of ion selectivity in channelrhodopsins. J. Vis. Exp. https://doi.org/10.3791/55497 (2017).
Dong, X.-P. et al. The type IV mucolipidosis-associated protein TRPML1 is an endolysosomal iron release channel. Nature 455, 992–996 (2008).
Surana, S., Bhat, J. M., Koushika, S. P. & Krishnan, Y. An autonomous DNA nanomachine maps spatiotemporal pH changes in a multicellular living organism. Nat. Commun. 2, 340 (2011).
Van Galen, J. et al. Sphingomyelin homeostasis is required to form functional enzymatic domains at the trans-Golgi network. J. Cell Biol. 206, 609–618 (2014).
Modi, S., Halder, S., Nizak, C. & Krishnan, Y. Recombinant antibody mediated delivery of organelle-specific DNA pH sensors along endocytic pathways. Nanoscale 6, 1144–1152 (2014).
Zhang, Q. et al. Functional relevance of Golgi- and plasma membrane-localized endothelial NO synthase in reconstituted endothelial cells. Arterioscler. Thromb. Vasc. Biol. 26, 1015–1021 (2006).
Acknowledgements
We thank J. Kuriyan, J. C. Clardy, J. W. Szostak, H. Xu and F. Bezanilla for critical comments; K. Chakraborty, X. Zhang, J. L. C. Souza, J. Delgado and Y. Jiang for discussions; and the integrated light microscopy and mass spectrometry facilities at the University of Chicago. This work was supported by the Women’s Board of the University of Chicago; FA9550-19-0003 from AFOSR, NIH grants R21NS114428, 1R01NS112139-01A1, Pilot and Feasibility award from an NIH-NIDDK Center grant P30DK42086 to the University of Chicago’s Digestive Diseases Research Center; and Chicago Biomedical Consortium with support from the Searle Funds at the Chicago Community Trust, C-084. M.S. was a Heisenberg fellow supported by the Deutsche Forschungsgemeinschaft.
Author information
Authors and Affiliations
Contributions
A.S., A.T.V. and Y.K. designed the project. A.S., A.T.V. and K.S.P. synthesized the probe, A.S. and J.D. built the instrumentation, and A.S., J.D. and B.S. performed the experiments. M.S. provided a scavenger receptor plasmid. A.S. and Y.K. designed the experiments and analysed and interpreted the data. A.S. and Y.K. wrote the paper. All authors provided input on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Nanotechnology thanks Yonggang Ke, Haoxing Xu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic of lysosomal Vmem measurement.
Imaging protocol of organelles labelled with VoltairIM. Resting organelles are imaged in the G and R channels, neutralized with valinomycin-monensin. The second set of images is acquired in the G and R channels. The resting membrane potential of the organelle is calculated by normalizing the G/R values from untreated lysosomes (xi) to neutralized lysosomes (yi).
Supplementary information
Supplementary Information
Supplementary Figs. 1–28, Notes 1–6, Tables 1,2.
Supplementary Video 1
Pseudo-coloured video showing the sensitivity of RVF. RVF-labelled HEK 293T cells were voltage clamped from −100 mV to +100 mV at increments of 10 mV in extracellular buffer. The clamped cell is shown by a white arrow; scale = 10 µm.
Supplementary Video 2
Pseudo-coloured video showing the sensitivity of VoltairPM. Labelled HEK 293T cells were voltage clamped from −100 mV to +100 mV at increments of 10 mV in extracellular buffer. The RVF channel shows the fluorescence change with respect to the membrane potential, whereas Atto647N fluorescence is insensitive to the applied voltage increments. The G/R ratio quantitatively shows the change in the membrane potential difference; scale = 10 µm.
Supplementary Video 3
Time-lapse imaging of lysosomal membrane potential in COS-7 cells labelled with VoltairIM. ML-SA1 (20 µM) or Vehicle (DMSO) is added to cells at 100 s. The pseudo-coloured images represent the computed intensity ratio of VoltairIM in the G (RVF) and R (Atto647N) channels. The cell of interest is represented by a white ROI (region of interest); scale = 10 µm.
Supplementary Video 4
Time-lapse imaging of endosomal membrane potential in COS-7 cells labelled with VoltairIM. ML-SA1 (20 µM) is added to cells at 100 s. The pseudo-coloured images represent the computed intensity ratio of VoltairIM in the G (RVF) and R (Atto647N) channels. The cell of interest is represented by a white ROI (region of interest); scale = 10 µm.
Supplementary Video 5
Time-lapse imaging of cytosolic calcium levels in ATP-treated COS-7 cells, labelled with Fluo-4 AM. 100 µM ATP is added to cells at 35 s. The pseudo-coloured images represent the intensity of Fluo-4 in a heat map; scale = 10 µm.
Supplementary Video 6
Time-lapse imaging of lysosomal membrane potential in ATP-treated COS-7 cells labelled with VoltairIM. 100 µM ATP or 1X PBS is added to cells at 35 s. The pseudo-coloured images represent the computed intensity ratio of VoltairIM in the G (RVF) and R (Atto647N) channels. The cell of interest is represented by a white ROI (region of interest). The white arrowheads indicate the lysosomes undergoing ATP-induced hyperpolarization. The decrease in G/R (observed in movie) represents the increase in positive membrane potential; scale = 10 µm.
Rights and permissions
About this article
Cite this article
Saminathan, A., Devany, J., Veetil, A.T. et al. A DNA-based voltmeter for organelles. Nat. Nanotechnol. 16, 96–103 (2021). https://doi.org/10.1038/s41565-020-00784-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41565-020-00784-1
This article is cited by
-
DNA-functionalized artificial mechanoreceptor for de novo force-responsive signaling
Nature Chemical Biology (2024)
-
Monitoring of Cell Membrane Microenvironment Based on DNA Nanodevices
Chemical Research in Chinese Universities (2024)
-
Detecting organelle-specific activity of potassium channels with a DNA nanodevice
Nature Biotechnology (2023)
-
DNA nanodevices map intracellular ions
Nature Biotechnology (2023)
-
A DNA nanodevice for mapping sodium at single-organelle resolution
Nature Biotechnology (2023)