Abstract
Background
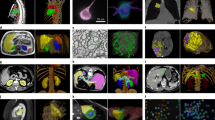
To validate the feasibility of building a deep learning model to predict axial length (AL) for moderate to high myopic patients from ultra-wide field (UWF) images.
Methods
This study included 6174 UWF images from 3134 myopic patients during 2014 to 2020 in Eye and ENT Hospital of Fudan University. Of 6174 images, 4939 were used for training, 617 for validation, and 618 for testing. The coefficient of determination (R2), mean absolute error (MAE), and mean squared error (MSE) were used for model performance evaluation.
Results
The model predicted AL with high accuracy. Evaluating performance of R2, MSE and MAE were 0.579, 1.419 and 0.9043, respectively. Prediction bias of 64.88% of the tests was under 1-mm error, 76.90% of tests was within the range of 5% error and 97.57% within 10% error. The prediction bias had a strong negative correlation with true AL values and showed significant difference between male and female (P < 0.001). Generated heatmaps demonstrated that the model focused on posterior atrophy changes in pathological fundus and peri-optic zone in normal fundus. In sex-specific models, R2, MSE, and MAE results of the female AL model were 0.411, 1.357, and 0.911 in female dataset and 0.343, 2.428, and 1.264 in male dataset. The corresponding metrics of male AL models were 0.216, 2.900, and 1.352 in male dataset and 0.083, 2.112, and 1.154 in female dataset.
Conclusions
It is feasible to utilize deep learning models to predict AL for moderate to high myopic patients with UWF images.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 18 print issues and online access
$259.00 per year
only $14.39 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Morgan IG, Ohno-Matsui K, Saw SM. Myopia. Lancet. 2012;379:1739–48.
Baird PN, Saw SM, Lanca C, Guggenheim JA, Smith Iii EL, Zhou X, et al. Myopia. Nat Rev Dis Prim. 2020;6:99.
Holden BA, Fricke TR, Wilson DA, Jong M, Naidoo KS, Sankaridurg P, et al. Global prevalence of myopia and high myopia and temporal trends from 2000 through 2050. Ophthalmology. 2016;123:1036–42.
Morgan IG, French AN, Ashby RS, Guo X, Ding X, He M, et al. The epidemics of myopia: Aetiology and prevention. Prog Retinal Eye Res. 2018;62:134–49.
Koh V, Tan C, Tan PT, Tan M, Balla V, Nah G, et al. Myopic maculopathy and optic disc changes in highly myopic young asian eyes and impact on visual acuity. Am J Ophthalmol. 2016;164:69–79.
Melles RB, Holladay JT, Chang WJ. Accuracy of intraocular lens calculation formulas. Ophthalmology. 2018;125:169–78.
Cen LP, Ji J, Lin JW, Ju ST, Lin HJ, Li TP, et al. Automatic detection of 39 fundus diseases and conditions in retinal photographs using deep neural networks. Nat Commun. 2021;12:4828.
Kim KM, Heo TY, Kim A, Kim J, Han KJ, Yun J, et al. Development of a fundus image-based deep learning diagnostic tool for various retinal diseases. JPM. 2021;11:321.
Grassmann F, Mengelkamp J, Brandl C, Harsch S, Zimmermann ME, Linkohr B, et al. A deep learning algorithm for prediction of age-related eye disease study severity scale for age-related macular degeneration from color fundus photography. Ophthalmology. 2018;125:1410–20.
Peng Y, Dharssi S, Chen Q, Keenan TD, Agrón E, Wong WT, et al. DeepSeeNet: a deep learning model for automated classification of patient-based age-related macular degeneration severity from color fundus photographs. Ophthalmology. 2019;126:565–75.
Kim KE, Kim JM, Song JE, Kee C, Han JC, Hyun SH. Development and validation of a deep learning system for diagnosing glaucoma using optical coherence tomography. JCM. 2020;9:2167.
Li Z, He Y, Keel S, Meng W, Chang RT, He M. Efficacy of a deep learning system for detecting glaucomatous optic neuropathy based on color fundus photographs. Ophthalmology. 2018;125:1199–206.
Li Z, Guo C, Lin D, Nie D, Zhu Y, Chen C, et al. Deep learning for automated glaucomatous optic neuropathy detection from ultra-widefield fundus images. Br J Ophthalmol. 2021;105:1548–54.
Fu H, Li F, Xu Y, Liao J, Xiong J, Shen J, et al. A retrospective comparison of deep learning to manual annotations for optic disc and optic cup segmentation in fundus photographs. Trans Vis Sci Tech. 2020;9:33.
Liefers B, Colijn JM, González-Gonzalo C, Verzijden T, Wang JJ, Joachim N, et al. A deep learning model for segmentation of geographic atrophy to study its long-term natural history. Ophthalmology. 2020;127:1086–96.
Maloca PM, Lee AY, De Carvalho ER, Okada M, Fasler K, Leung I, et al. Validation of automated artificial intelligence segmentation of optical coherence tomography images. Pławiak P, editor. PLoS ONE. 2019;14:e0220063.
Pekala M, Joshi N, Liu TYA, Bressler NM, DeBuc DC, Burlina P. Deep learning based retinal OCT segmentation. Comput Biol Med. 2019;114:103445.
Dong L, Hu XY, Yan YN, Zhang Q, Zhou N, Shao L, et al. Deep learning-based estimation of axial length and subfoveal choroidal thickness from color fundus photographs. Front Cell Dev Biol. 2021;9:653692.
Jeong Y, Lee B, Han JH, Oh J. Ocular axial length prediction based on visual interpretation of retinal fundus images via deep neural network. IEEE J Sel Top Quantum Electron. 2021;27:1–7.
Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z. Rethinking the Inception Architecture for Computer Vision. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 2016, pp. 2818–26, https://doi.org/10.1109/CVPR.2016.308.
He K, Zhang X, Ren S, Sun J. Delving Deep into Rectifiers: Surpassing Human-Level Performance on ImageNet Classification. 2015 IEEE International Conference on Computer Vision (ICCV), Santiago, Chile, 2015, pp. 1026–34, https://doi.org/10.1109/ICCV.2015.123.
Aravkin AY, Kambadur A, Lozano AC, Luss R. Sparse Quantile Huber Regression for Efficient and Robust Estimation. arXiv; 2014. Available from: http://arxiv.org/abs/1402.4624.
Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-CAM: visual explanations from deep networks via gradient-based localization. Int J Comput Vis. 2020;128:336–59.
Gu J, Oelke D. Understanding Bias in Machine Learning. arXiv; 2019. Available from: http://arxiv.org/abs/1909.01866.
Atchison DA, Pritchard N, Schmid KL, Scott DH, Jones CE, Pope JM. Shape of the retinal surface in emmetropia and myopia. Invest Ophthalmol Vis Sci. 2005;46:2698.
Hashimoto S, Yasuda M, Fujiwara K, Ueda E, Hata J, Hirakawa Y, et al. Association between axial length and myopic maculopathy. Ophthalmol Retin. 2019;3:867–73.
Ruiz-Medrano J, Montero JA, Flores-Moreno I, Arias L, García-Layana A, Ruiz-Moreno JM. Myopic maculopathy: current status and proposal for a new classification and grading system (ATN). Prog Retinal Eye Res. 2019;69:80–115.
Ohno-Matsui K, Lai TYY, Lai CC, Cheung CMG. Updates of pathologic myopia. Prog Retinal Eye Res. 2016;52:156–87.
Guo X, Li R, Lu X, Zhang X, Wu Q, Tian Q, et al. Quantization of optic disc characteristics in young adults based on artificial intelligence. Curr Eye Res. 20239;1–10.
Qiao Y, Cheng D, Zhu X, Ruan K, Ye Y, Yu J, et al. Characteristics of the peripapillary structure and vasculature in patients with myopic anisometropia. Trans Vis Sci Tech. 2023;12:16.
Cheng D, Ruan K, Wu M, Qiao Y, Gao W, Lian H, et al. Characteristics of the optic nerve head in myopic eyes using swept-source optical coherence tomography. Invest Ophthalmol Vis Sci. 2022;63:20.
He J, Ye L, Chu C, Chen Q, Sun D, Xie J, et al. Using a combination of peripapillary atrophy area and choroidal thickness for the prediction of different types of myopic maculopathy. Eye (Lond). 2023;37:2801–9.
Acknowledgements
The authors appreciated all the colleagues involved in the whole process of the research.
Funding
1) National Natural Science Foundation of China (Grant No. 82301251). 2) Joint research project of new frontier technology in municipal hospitals (SHDC12018103). 3) Project of Shanghai Science and Technology (Grant No.20410710100). 4) Clinical Research Plan of SHDC (SHDC2020CR1043B). 5) Project of Shanghai Xuhui District Science and Technology (2020-015). 6) Shanghai Rising-Star Program (21QA1401500). 7) Shanghai Yangfan Project (23YF1445300).
Author information
Authors and Affiliations
Contributions
YW: conceptualization, formal analysis, writing original draft; RW, DY: data acquisition, formal analysis, visualization; KS: model development; YS, LN: review and editing; ML, XZ: conceptualization, supervision, funding acquisition.
Corresponding authors
Ethics declarations
competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, Y., Wei, R., Yang, D. et al. Development and validation of a deep learning model to predict axial length from ultra-wide field images. Eye 38, 1296–1300 (2024). https://doi.org/10.1038/s41433-023-02885-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41433-023-02885-2