Abstract
Background
Gut-derived metabolites, products of microbial and host co-metabolism, may inform mechanisms underlying children’s neurodevelopment. We investigated whether infant fecal metabolites were related to toddler social behavior.
Methods
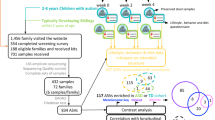
Stool samples collected from 6-week-olds (n = 86) and 1-year-olds (n = 209) in the New Hampshire Birth Cohort Study (NHBCS) were analyzed using nuclear magnetic resonance spectroscopy metabolomics. Autism-related behavior in 3-year-olds was assessed by caregivers using the Social Responsiveness Scale (SRS-2). To assess the association between metabolites and SRS-2 scores, we used a traditional single-metabolite approach, quantitative metabolite set enrichment (QEA), and self-organizing maps (SOMs).
Results
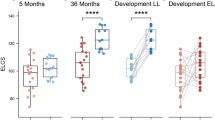
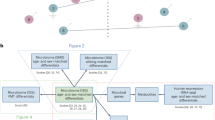
Using a single-metabolite approach and QEA, no individual fecal metabolite or metabolite set at either age was associated with SRS-2 scores. Using the SOM method, fecal metabolites of six-week-olds organized into four profiles, which were unrelated to SRS-2 scores. In 1-year-olds, one of twelve fecal metabolite profiles was associated with fewer autism-related behaviors, with SRS-2 scores 3.4 (95%CI: −7, 0.2) points lower than the referent group. This profile had higher concentrations of lactate and lower concentrations of short chain fatty acids than the reference.
Conclusions
We uncovered metabolic profiles in infant stool associated with subsequent social behavior, highlighting one potential mechanism by which gut bacteria may influence neurobehavior.
Impact
-
Differences in host and microbial metabolism may explain variability in neurobehavioral phenotypes, but prior studies do not have consistent results.
-
We applied three statistical techniques to explore fecal metabolite differences related to social behavior, including self-organizing maps (SOMs), a novel machine learning algorithm.
-
A 1-year-old fecal metabolite pattern characterized by high lactate and low short-chain fatty acid concentrations, identified using SOMs, was associated with social behavior less indicative of autism spectrum disorder.
-
Our findings suggest that social behavior may be related to metabolite profiles and that future studies may uncover novel findings by applying the SOM algorithm.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 14 print issues and online access
$259.00 per year
only $18.50 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
The datasets analyzed during the current study may be made available from the corresponding author on reasonable request.
References
Thomson, S. et al. Increased maternal non-oxidative energy metabolism mediates association between prenatal di-(2-ethylhexyl) phthalate (DEHP) exposure and offspring autism spectrum disorder symptoms in early life: a birth cohort study. Environ. Int. 171, 107678 (2023).
Gagliano, A. et al. 1H-NMR-based metabolomics in autism spectrum disorder and pediatric acute-onset neuropsychiatric syndrome. J. Clin. Med 11, 6493 (2022).
Galineau, L. et al. Glutamatergic synapse in autism: a complex story for a complex disorder. Mol. Psychiatry https://doi.org/10.1038/s41380-022-01860-9 (2022).
Brister, D. et al. Metabolomic signatures of autism spectrum disorder. J. Pers. Med. 12, 1727 (2022).
Parenti, M. et al. Maternal serum and placental metabolomes in association with prenatal phthalate exposure and neurodevelopmental outcomes in the MARBLES cohort. Metabolites 12, 829 (2022).
Jones, J., Reinke, S. N., Mousavi-Derazmahalleh, M., Palmer, D. J. & Christophersen, C. T. Changes to the gut microbiome in young children showing early behavioral signs of autism. Front. Microbiol. 13, 905901 (2022).
Likhitweerawong, N. et al. Profiles of urine and blood metabolomics in autism spectrum disorders. Metab. Brain Dis. 36, 1641–1671 (2021).
Deng, K. et al. Comparison of fecal and blood metabolome reveals inconsistent associations of the gut microbiota with cardiometabolic diseases. Nat. Commun. 14, 571 (2023).
Wishart, D. S. et al. NMR and metabolomics—a roadmap for the future. Metabolites 12, 678 (2022).
Kim, S. J., Song, H. E., Lee, H. Y. & Yoo, H. J. Mass spectrometry-based metabolomics in translational research. in Advanced Imaging and Bio Techniques for Convergence Science (eds Kim, J. K., Kim, J. K. & Pack, C.-G.) 509–531 (Springer, 2021). https://doi.org/10.1007/978-981-33-6064-8_19.
Pang, Z. et al. MetaboAnalyst 5.0: narrowing the gap between raw spectra and functional insights. Nucleic Acids Res. 49, W388–W396 (2021).
Pearce, J. L. et al. Characterizing the spatial distribution of multiple pollutants and populations at risk In Atlanta, Georgia. Spat. Spatiotemporal Epidemiol. 18, 13–23 (2016).
Pearce, J. L. et al. Using self-organizing maps to develop ambient air quality classifications: a time series example. Environ. Health 13, 56 (2014).
Gilbert-Diamond, D. et al. Rice consumption contributes to arsenic exposure in US women. Proc. Natl. Acad. Sci. USA 108, 20656–20660 (2011).
Madan, J. C. et al. Association of cesarean delivery and formula supplementation with the intestinal microbiome of 6-week-old infants. JAMA Pediatr. 170, 212–219 (2016).
Karagas, M. R. et al. Alterations in microbial-associated fecal metabolites in relation to arsenic exposure among infants. Expo Health 1–9 https://doi.org/10.1007/s12403-022-00468-2 (2022).
Hoen, A. G. et al. Association of Cesarean delivery and formula supplementation with the stool metabolome of 6-week-old infants. Metabolites 11, 702 (2021).
Li, J. V. et al. Global metabolic responses of NMRI mice to an experimental Plasmodium berghei infection. J. Proteome Res. 7, 3948–3956 (2008).
Zheng, X. et al. The footprints of gut microbial-mammalian co-metabolism. J. Proteome Res. 10, 5512–5522 (2011).
Nicholson, J. K. et al. Metabolic phenotyping in clinical and surgical environments. Nature 491, 384–392 (2012).
Weljie, A. M., Newton, J., Mercier, P., Carlson, E. & Slupsky, C. M. Targeted profiling: quantitative analysis of 1H NMR metabolomics data. Anal. Chem. 78, 4430–4442 (2006).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Constantino, J. N. & Gruber, C. P. Social Responsiveness Scale—Second Edition (SRS-2) (Western Psychological Services, 2012).
Bruni, T. P. Test review: social responsiveness scale–second edition (SRS-2). J. Psychoeduc. Assess. 32, 365–369 (2014).
Azur, M. J., Stuart, E. A., Frangakis, C. & Leaf, P. J. Multiple imputation by chained equations: what is it and how does it work? Int. J. Methods Psychiatr. Res. 20, 40–49 (2011).
van Buuren, S. & Groothuis-Oudshoorn, K. mice: multivariate imputation by chained equations in R. J. Stat. Softw. 45, 1–67 (2011).
Harel, O. et al. Multiple imputation for incomplete data in epidemiologic studies. Am. J. Epidemiol. 187, 576–584 (2018).
Rubin, D. B. Multiple Imputation for Nonresponse in Surveys (John Wiley & Sons, 2004).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B (Methodol.) 57, 289–300 (1995).
Højsgaard, S., Halekoh, U. & Yan, J. The R package geepack for generalized estimating equations. J. Stat. Softw. 15, 1–11 (2006).
Xia, J. & Wishart, D. S. MSEA: a web-based tool to identify biologically meaningful patterns in quantitative metabolomic data. Nucleic Acids Res. 38, W71–W77 (2010).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 44, D457–D462 (2016).
Kanehisa, M., Furumichi, M., Sato, Y., Ishiguro-Watanabe, M. & Tanabe, M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 49, D545–D551 (2021).
Yao, Q. et al. Global prioritization of disease candidate metabolites based on a multi-omics composite network. Sci. Rep. 5, 17201 (2015).
Laaksonen, J. & Honkela, T. (eds) Advances in Self-Organizing Maps. In Proc. 8th International Workshop, WSOM 2011 (Springer, 2011).
Kohonen, T. Essentials of the self-organizing map. Neural Netw. 37, 52–65 (2013).
Doherty, B. T., Pearce, J. L., Anderson, K. A., Karagas, M. R. & Romano, M. E. Assessment of multipollutant exposures during pregnancy using silicone wristbands. Front Public Health 8, 547239 (2020).
Laue, H. E., Coker, M. O. & Madan, J. C. The developing microbiome from birth to 3 years: the gut-brain axis and neurodevelopmental outcomes. Front. Pediatr. 10, 815885 (2022).
Needham, B. D. et al. Plasma and fecal metabolite profiles in autism spectrum disorder. Biol. Psychiatry 89, 451–462 (2021).
De Angelis, M. et al. Fecal microbiota and metabolome of children with autism and pervasive developmental disorder not otherwise specified. PLoS One 8, e76993 (2013).
Kang, D.-W. et al. Differences in fecal microbial metabolites and microbiota of children with autism spectrum disorders. Anaerobe 49, 121–131 (2018).
Wang, M. et al. Alteration of gut microbiota-associated epitopes in children with autism spectrum disorders. Brain Behav. Immun. 75, 192–199 (2019).
Zhu, J. et al. Alterations in gut vitamin and amino acid metabolism are associated with symptoms and neurodevelopment in children with autism spectrum disorder. J. Autism Dev. Disord. 52, 3116–3128 (2022).
Yap, C. X. et al. Autism-related dietary preferences mediate autism-gut microbiome associations. Cell 184, 5916–5931.e17 (2021).
Sharon, G. et al. Human gut microbiota from autism spectrum disorder promote behavioral symptoms in mice. Cell 177, 1600–1618.e17 (2019).
Dan, Z. et al. Altered gut microbial profile is associated with abnormal metabolism activity of Autism Spectrum Disorder. Gut Microbes 11, 1246–1267 (2020).
Muller, C. L., Anacker, A. M. J. & Veenstra-VanderWeele, J. The serotonin system in autism spectrum disorder: from biomarker to animal models. Neuroscience 321, 24–41 (2016).
Wang, S. P. et al. Pivotal roles for pH, lactate, and lactate-utilizing bacteria in the stability of a human colonic microbial ecosystem. mSystems 5, e00645–20 (2020).
Ho, L. K. H. et al. Gut microbiota changes in children with autism spectrum disorder: a systematic review. Gut Pathog. 12, 6 (2020).
Laue, H. E., Korrick, S. A., Baker, E. R., Karagas, M. R. & Madan, J. C. Prospective associations of the infant gut microbiome and microbial function with social behaviors related to autism at age 3 years. Sci. Rep. 10, 15515 (2020).
Wyss, M. T., Jolivet, R., Buck, A., Magistretti, P. J. & Weber, B. In vivo evidence for lactate as a neuronal energy source. J. Neurosci. 31, 7477–7485 (2011).
Holmgren, C. D. et al. Energy substrate availability as a determinant of neuronal resting potential, GABA signaling and spontaneous network activity in the neonatal cortex in vitro. J. Neurochem. 112, 900–912 (2010).
Coretti, L. et al. Gut microbiota features in young children with autism spectrum disorders. Front. Microbiol. 9, 3146 (2018).
Rosen, N. J., Yoshida, C. K. & Croen, L. A. Infection in the first 2 years of life and autism spectrum disorders. Pediatrics 119, e61–e69 (2007).
Chalifour, B. et al. The potential role of early life feeding patterns in shaping the infant fecal metabolome: implications for neurodevelopmental outcomes. npj Metab. Health Dis. 1, 1–13 (2023).
Holzhausen, E. A. et al. Longitudinal profiles of the fecal metabolome during the first 2 years of life. Sci. Rep. 13, 1886 (2023).
McKeen, S. et al. Adaptation of the infant gut microbiome during the complementary feeding transition. Plos One 17, e0270213 (2022).
Funding
This work was supported by the National Institutes of Health Office of the Director (UH3OD023275), the National Institute of Environmental Health Science (K99ES034086, P01ES022832, P42ES007373), the National Institute of General Medical Sciences (P20GM104416), the National Institute of Diabetes and Digestive and Kidney Diseases (U24DK097193), the National Cancer Institute (T32CA134286), and the U.S. Environmental Protection Agency (RD-83544201). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health or the Environmental Protection Agency.
Author information
Authors and Affiliations
Contributions
All authors meet the authorship requirements and approve the final version to be published. H.E.L.: conceptualization, methodology, formal analysis, writing – original draft, writing – review and editing, visualization, funding acquisition; J.A.B.: conceptualization, methodology, writing - review and editing; S.S.: investigation, resources, data curation, writing – review and editing; S.M.: investigation, resources, data curation, writing – review and editing; W.P.: investigation, resources, data curation, writing – review and editing; T.J.P.: investigation, resources, data curation, writing – review and editing; A.G.H.: resources, writing - review and editing, funding acquisition; J.C.M.: resources, writing - review and editing, funding acquisition; M.R.K.: conceptualization, methodology, resources, writing – review and editing, project administration, funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Caregivers of participants provided written informed consent. All study protocols were approved by the Center for the Protection of Human Subjects at Dartmouth, and all methods were carried out in accordance with relevant guidelines and regulations.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Laue, H.E., Bauer, J.A., Pathmasiri, W. et al. Patterns of infant fecal metabolite concentrations and social behavioral development in toddlers. Pediatr Res (2024). https://doi.org/10.1038/s41390-024-03129-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41390-024-03129-z