Abstract
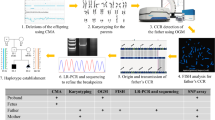
Chromosomal rearrangements mostly result from non-allelic homologous recombination mediated by low-copy repeats (LCRs) or segmental duplications (SDs). Recent studies on recombinant chromosome 18 (rec (18)) have focused on diagnoses and clinical phenotypes. We diagnosed two cases of prenatal rec (18) and identified precise breakpoint intervals using karyotype and chromosomal microarray analyses. We analyzed the distribution characteristics of breakpoint repetitive elements to infer rearrangement mechanisms and reviewed relevant literature to identify genetic trends. Among the 12 families with 25 pregnancies analyzed, 68% rec (18), 24% spontaneous abortions, and 8% normal births were reported. In the 17 rec (18) cases, 65% presented maternal origin and 35% were paternal. Short-arm breakpoints at p11.31 were reported in 10 cases, whereas the long-arm breakpoints were located at q21.3 (6 cases) and q12 (4 cases). Breakpoints of pericentric inversions on chromosome 18 are concentrated in p11.31, q21.3, and q12 regions. Rearrangements at 18p11.31 are non-recurrent events. ALUs, LINE1s, and MIRs were enriched at the breakpoint regions (1.85 to 3.42-fold enrichment over the entire chromosome 18), while SDs and LCRs were absent. ALU subfamilies had sequence identities of 85.94% and 83.01% between two pair breakpoints. Small repetitive elements may mediate recombination-coupled DNA repair processes, facilitating rearrangements on chromosome 18. Maternal inversion carriers are more prone to abnormal recombination in prenatal families with rec (18). Recombinant chromosomes may present preferential segregation during gamete formation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All relevant data of this study are presented.
References
Martin AO, Simpson JL, Deddish RB, Elias S. Clinical implications of chromosomal inversions. A pericentric inversion in No. 18 segregating in a family ascertained through an abnormal proband. Am J Perinatol. 1983;1:81–8.
Sujansky E, Smith AC, Peakman DC, McConnell TS, Baca P, Robinson A. Familial pericentric inversion of chromosome 8. Am J Med Genet. 1981;10:229–35.
Lee JA, Carvalho CM, Lupski JR. A DNA replication mechanism for generating nonrecurrent rearrangements associated with genomic disorders. Cell. 2007;131:1235–47.
Weckselblatt B, Rudd MK. Human structural variation: mechanisms of chromosome rearrangements. Trends Genet. 2015;31:587–99.
Samonte RV, Eichler EE. Segmental duplications and the evolution of the primate genome. Nat Rev Genet. 2002;3:65–72.
Abdullaev E. Dynamical aspects of the evolution of segmental duplications in the human genome. Berlin: Freie Universität Berlin; 2022. p. 3–33.
Kojima KK. LINEs contribute to the origins of middle bodies of SINEs besides 3’ tails. Genome Biol Evol. 2018;10:370–79.
Lee MJ, Park SH, Shim SH, Moon MJ, Cha DH. Prenatal diagnosis and molecular cytogenetic characterization of partial dup(18q)/del(18p) due to a paternal pericentric inversion 18 in a fetus with multiple anomalies. Taiwan J Obstet Gynecol. 2019;58:318–23.
Yu J, Xiaolu C, Jiayan C, Meijiao C, Jian Z, Yunsheng G, et al. Prenatal diagnosis and genetic counseling for two pedigrees with pericentric inversion of chromosome 18. Chin J Perinat Med. 2019;22:127–33.
Shuang C, Dejun L, Chunzhu J, Linlin L. Prenatal diagnosis of a case of 46, XY, rec(18)dup(18q)inv(18)(p11q12). Chin J Med Genet. 2019;36:681–81.
Zamani AG, Acar A, Durakbasi-Dursun G, Yildirim MS, Ceylaner S, Tuncez E. Recurrent proximal 18p monosomy and 18q trisomy in a family due to a pericentric inversion. Am J Med Genet A. 2014;164a:1239–44.
Sahin FI, Ozer O, Tarim E, Yilmaz Z. Detection of identical unbalanced karyotype in two consequent fetuses due to a maternal pericentric inversion of chromosome 18. J Obstet Gynaecol. 2012;32:698–700.
Kariminejad A, Kariminejad R, Moshtagh A, Zanganeh M, Kariminejad MH, Neuenschwander S, et al. Pericentric inversion of chromosome 18 in parents leading to a phenotypically normal child with segmental uniparental disomy 18. Eur J Hum Genet. 2011;19:555–60.
Prabhakara K, Wyandt HE, Huang XL, Prasad KS, Ramadevi AR. Recurrent proximal 18p monosomy and 18q trisomy in a family with a maternal pericentric inversion of chromosome 18. Ann Genet. 2004;47:297–303.
Roberts D, Sweeney E, Walkinshaw S. Congenital cystic adenomatoid malformation of the lung coexisting with recombinant chromosome 18. A case report. Fetal Diagn Ther. 2001;16:65–7.
Leonard NJ, Tomkins DJ, Demianczuk N. Prenatal diagnosis of holoprosencephaly (HPE) in a fetus with a recombinant (18)dup(18q)inv(18)(p11.31q11.2)mat. Prenat Diagn. 2000;20:947–9.
Mohsen-Pour N, Talebi T, Naderi N, Moghadam MH, Maleki M, Kalayinia S. Chromosome 9 Inversion: Pathogenic or Benign? A comprehensive systematic review of all clinical reports. Curr Mol Med. 2022;22:385–400.
Stapley J, Feulner PGD, Johnston SE, Santure AW, Smadja CM. Variation in recombination frequency and distribution across eukaryotes: patterns and processes. Philos Trans R Soc Lond B Biol Sci. 2017;372:20160455.
Liehr T, Weise A, Mrasek K, Ziegler M, Padutsch N, Wilhelm K, et al. Recombinant chromosomes resulting from parental pericentric inversions-two new cases and a review of the literature. Front Genet. 2019;10:1165.
Bailey JA, Gu Z, Clark RA, Reinert K, Samonte RV, Schwartz S, et al. Recent segmental duplications in the human genome. Science. 2002;297:1003–7.
Nusbaum C, Zody MC, Borowsky ML, Kamal M, Kodira CD, Taylor TD, et al. DNA sequence and analysis of human chromosome 18. Nature. 2005;437:551–5.
Startek M, Szafranski P, Gambin T, Campbell IM, Hixson P, Shaw CA, et al. Genome-wide analyses of LINE-LINE-mediated nonallelic homologous recombination. Nucleic Acids Res. 2015;43:2188–98.
Rossetti LC, Goodeve A, Larripa IB, De Brasi CD. Homeologous recombination between AluSx-sequences as a cause of hemophilia. Hum Mutat. 2004;24:440.
Weckselblatt B, Hermetz KE, Rudd MK. Unbalanced translocations arise from diverse mutational mechanisms including chromothripsis. Genome Res. 2015;25:937–47.
Gu S, Yuan B, Campbell IM, Beck CR, Carvalho CM, Nagamani SC, et al. Alu-mediated diverse and complex pathogenic copy-number variants within human chromosome 17 at p13.3. Hum Mol Genet. 2015;24:4061–77.
Carvalho CM, Lupski JR. Mechanisms underlying structural variant formation in genomic disorders. Nat Rev Genet. 2016;17:224–38.
Vianna-Morgante AM, Nozaki MJ, Ortega CC, Coates V, Yamamura Y. Partial monosomy and partial trisomy 18 in two offspring of carrier of pericentric inversion of chromosome 18. J Med Genet. 1976;13:366–70.
Anton E, Vidal F, Egozcue J, Blanco J. Genetic reproductive risk in inversion carriers. Fertil Steril. 2006;85:661–6.
Morel F, Laudier B, Guérif F, Couet ML, Royère D, Roux C, et al. Meiotic segregation analysis in spermatozoa of pericentric inversion carriers using fluorescence in-situ hybridization. Hum Reprod. 2007;22:136–41.
Asano T, Ikeuchi T, Shinohara T, Enokido H, Hashimoto K. Partial 18q trisomy and 18p monosomy resulting from a maternal pericentric inversion, inv(18)(p11.2q21.3). Jinrui Idengaku Zasshi. 1991;36:257–65.
Gardner RM, Sutherland GR, Shaffer LG. Chromosome abnormalities and genetic counseling. USA: OUP; 2011.
Turleau C. Monosomy 18p. Orphanet J Rare Dis. 2008;3:4.
Wester U, Bondeson ML, Edeby C, Annerén G. Clinical and molecular characterization of individuals with 18p deletion: a genotype-phenotype correlation. Am J Med Genet A. 2006;140:1164–71.
Prontera P, Buldrini B, Aiello V, Rogaia D, Mencarelli A, Gruppioni R, et al. Familial pericentric inversion of chromosome 18: intrafamilial variability of the recombinant dup(18q). Genet Couns. 2010;21:91–7.
Vermeulen SJ, Speleman F, Vanransbeeck L, Verspeet J, Menten B, Verschraegen-Spae MR, et al. Familial pericentric inversion of chromosome 18: behavioral abnormalities in patients heterozygous for either the dup(18p)/del(18q) or dup(18q)/del(18p) recombinant chromosome. Eur J Hum Genet. 2005;13:52–8.
Poterico JA, Vásquez F, Chávez-Pastor M, Trubnykova M, Chavesta F, Chirinos J, et al. A peruvian child with 18p-/18q+ syndrome and persistent microscopic hematuria. J Pediatr Genet. 2017;6:258–66.
Liehr T, Kosayakova N, Schröder J, Ziegler M, Kreskowski K, Pohle B, et al. Evidence for correlation of fragile sites and chromosomal breakpoints in carriers of constitutional balanced chromosomal rearrangements. Balk J Med Genet. 2011;14:13–6.
Durkin SG, Glover TW. Chromosome fragile sites. Annu Rev Genet. 2007;41:169–92.
Acknowledgements
All authors have contributed to the work, agree with the presented findings, and the work has not been published before nor is being considered for publication in another journal. We appreciate the contributions of our lab colleagues. Apologies to those whose work we are unable to cite. This research received no external funding.
Author information
Authors and Affiliations
Contributions
Design, conceptualization and supervision: LW and BD; methodology: YW, YX, HK, and LW; literature collection, bibliographic search and data curation: LW, BD, and YW; writing—original draft preparation: LW; writing—review and editing: LW; figures, table preparation and proofreading: LW and BD; All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study was conducted in accordance with the Declaration of Helsinki, and approved by Ethics Committee of Chengdu Women’s and Children’s Center Hospital(B2021(19) and Aug. 2021)” for studies involving humans.
Informed consent
Informed consent was obtained from all subjects involved in the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, L., Dong, B., Xie, Y. et al. The molecular mechanisms of recombinant chromosome 18 with parental pericentric inversions and a review of the literature. J Hum Genet 68, 625–634 (2023). https://doi.org/10.1038/s10038-023-01157-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-023-01157-x