Abstract
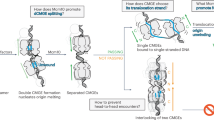
In bacteria, the processing of double-strand DNA breaks is mediated by the RecBCD, AddAB and AdnAB complexes. These multisubunit helicase–nuclease machines resect the DNA ends and load RecA protein to initiate homologous recombination. Recent studies have revealed fascinating insights into the molecular mechanisms of this process and the evolution of these machines.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chapman, J. R., Taylor, M. R. & Boulton, S. J. Playing the end game: DNA double-strand break repair pathway choice. Mol. Cell 47, 497–510 (2012).
Lieber, M. R. The mechanism of double-strand DNA break repair by the nonhomologous DNA end-joining pathway. Annu. Rev. Biochem. 79, 181–211 (2010).
Shuman, S. & Glickman, M. S. Bacterial DNA repair by non-homologous end joining. Nature Rev. Microbiol. 5, 852–861 (2007).
Chayot, R., Montagne, B., Mazel, D. & Ricchetti, M. An end-joining repair mechanism in Escherichia coli. Proc. Natl Acad. Sci. USA 107, 2141–2146 (2010).
Ayora, S. et al. Double-strand break repair in bacteria: a view from Bacillus subtilis. FEMS Microbiol. Rev. 35, 1055–1081 (2011).
Singleton, M. R., Dillingham, M. S., Gaudier, M., Kowalczykowski, S. C. & Wigley, D. B. Crystal structure of RecBCD enzyme reveals a machine for processing DNA breaks. Nature 432, 187–193 (2004).
Singleton, M. R., Dillingham, M. S. & Wigley, D. B. Structure and mechanism of helicases and nucleic acid translocases. Ann. Rev. Biochem. 76, 23–50 (2007).
Spies, M. & Kowalczykowski, S. C. The RecA binding locus of RecBCD is a general domain for recruitment of DNA strand exchange proteins. Mol. Cell 21, 573–580 (2006).
Taylor, A. F. & Smith, G. R. RecBC enzyme is a DNA helicase with fast and slow motors of opposite polarity. Nature 423, 869–893 (2003).
Dillingham, M. S., Spies, M. & Kowalczykowski, S. C. RecBCD enzyme is a bipolar DNA helicase. Nature 423, 893–897 (2003).
Dixon, D. A. & Kowalczykowski, S. C. The recombination hotspot chi is a regulatory sequence that acts by attenuating the nuclease activity of the E. coli RecBCD enzyme. Cell 73, 87–96 (1993).
Bianco, P. R. & Kowalczykowski, S. C. The recombination hot spot Chi is recognized by the translocating RecBCD enzyme as the single strand of DNA containing the sequence 5′-GCTGGTGG-3′. Proc. Natl Acad. Sci. USA 94, 6706–6711 (1997).
Anderson, D. G. & Kowalczykowski, S. C. The translocating RecBCD enzyme stimulates recombination by directing RecA protein onto ssDNA in a χ-regulated manner. Cell 90, 77–86 (1997).
Saikrishnan, K., Griffiths, S. P., Cook, N., Court, R. & Wigley, D. B. DNA binding to RecD: role of the 1B domain in SF1B helicase activity. EMBO J. 27, 2222–2229 (2008).
Velankar, S. S., Soultanas, P., Dillingham, M. S., Subramanya, H. S. & Wigley, D. B. Crystal structures of complexes of PcrA helicase with a DNA substrate indicate an inchworm mechanism. Cell 97, 75–84 (1999).
Saikrishnan, K., Powell, B., Cook, N. J., Webb, M. R. & Wigley, D. B. Mechanistic basis of 5′–3′ translocation in SF1B helicases. Cell 137, 849–859 (2009).
Smith, G. R. How RecBCD enzyme and Chi promote DNA break repair and recombination: a molecular biologist's view. Microbiol. Mol. Biol. Rev. 76, 217–228 (2012).
Farah, J. A. & Smith, G. R. The RecBCD enzyme initiation complex for DNA unwinding: enzyme positioning and DNA opening. J. Mol. Biol. 272, 699–715 (1997).
Wu, C. G., Bradford, C. & Lohman, T. M. Escherichia coli RecBC helicase has two translocase activities controlled by a single ATPase motor. Nature Struct. Mol. Biol. 17, 1210–1217 (2010).
Wu, C. G., Xie, F. & Lohman, T. M. The primary and secondary translocase activities within E. coli RecBC helicase are tightly coupled to ATP hydrolysis by the RecB motor. J. Mol. Biol. 423, 303–314 (2012).
Spies, M. et al. A molecular throttle: the recombination hotspot chi controls DNA translocation by the RecBCD helicase. Cell 114, 647–654 (2003).
Spies, M., Amitani, I., Baskin, R. J. & Kowalczykowski, S. C. RecBCD enzyme switches lead motor subunits in response to χ recognition. Cell 131, 694–705 (2007).
Handa, N., Bianco, P. R., Baskin, R. J. & Kowalczykowski, S. C. Direct visualization of RecBCD movement reveals cotranslocation of the RecD motor after chi recognition. Mol. Cell 17, 745–750 (2005).
Handa, N. et al. Molecular determinants responsible for recognition of the single-stranded DNA regulatory sequence, χ, by RecBCD enzyme. Proc. Natl Acad. Sci. USA 109, 8901–8906 (2012).
Yang, L. et al. Alteration of χ recognition by RecBCD reveals a regulated molecular latch and suggests a channel-bypass mechanism for biological control. Proc. Natl Acad. Sci. USA 109, 8907–8912 (2012).
Chédin, F., Noirot, P., Biaudet, V. & Ehrlich, S. D. A five-nucleotide sequence protects DNA from exonucleolytic degradation by AddAB, the RecBCD analogue of Bacillus subtilis. Mol. Microbiol. 29, 1369–1377 (1998).
Chédin, F., Handa, N., Dillingham, M. S. & Kowalczykowski, S. C. The AddAB helicase/nuclease forms a stable complex with its cognate χ sequence during translocation. J. Biol. Chem. 281, 18610–18617 (2006).
Saikrishnan, K. et al. Insights into Chi recognition from the structure of an AddAB-type helicase–nuclease complex. EMBO J. 31, 1568–1578 (2012).
Yeeles, J. T., Gwynn, E. J., Webb, M. R. & Dillingham, M. S. The AddAB helicase-nuclease catalyses rapid and processive DNA unwinding using a single Superfamily 1A motor domain. Nucleic Acids Res. 39, 2271–2285 (2011).
Yeeles, J. T. & Dillingham, M. S. A dual-nuclease mechanism for DNA break processing by AddAB-type helicase-nucleases. J. Mol. Biol. 371, 66–78 (2007).
Yeeles, J. T., van Aelst, K., Dillingham, M. S. & Moreno-Herrero, F. Recombination hotspots and single-stranded DNA binding proteins couple DNA translocation to DNA unwinding by the AddAB helicase-nuclease. Mol. Cell 42, 806–816 (2011).
Sinha, K. M., Unciuleac, M. C., Glickman, M. S. & Shuman, S. AdnAB: a new DSB-resecting motor–nuclease from mycobacteria. Genes Dev. 23, 1423–1437 (2009).
Unciuleac, M. C. & Shuman, S. Characterization of the mycobacterial AdnAB DNA motor provides insights into the evolution of bacterial motor-nuclease machines. J. Biol. Chem. 285, 2632–2641 (2010).
Unciuleac, M. C. & Shuman, S. Double strand break unwinding and resection by the mycobacterial helicase-nuclease AdnAB in the presence of single strand DNA-binding protein (SSB). J. Biol. Chem. 285, 34319–34329 (2010).
Lam, S. T., Stahl, M. M., McMilin, K. D. & Stahl, F. W. Rec-mediated recombinational hot spot activity in bacteriophage lambda. II. A mutation which causes hot spot activity. Genetics 77, 425–433 (1974).
Halpern, D. et al. Identification of DNA motifs implicated in maintenance of bacterial core genomes by predictive modeling. PLoS Genet. 3, 1614–1621 (2007).
Dixon, D. A., Churchill, J. J. & Kowalczykowski, S. C. Reversible inactivation of the Escherichia coli RecBCD enzyme by the recombination hotspot χ in vitro: evidence for functional inactivation or loss of the RecD subunit. Proc. Natl Acad. Sci. USA 91, 2980–2984 (1994).
Korolev, S., Hsieh, J., Gauss, G. H., Lohman, T. M. & Waksman, G. Major domain swiveling revealed by the crystal structures of complexes of E. coli Rep helicase bound to single-stranded DNA and ADP. Cell 90, 635–647 (1997).
Acknowledgements
D.B.W. is funded by Cancer Research UK and the Wellcome Trust.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Rights and permissions
About this article
Cite this article
Wigley, D. Bacterial DNA repair: recent insights into the mechanism of RecBCD, AddAB and AdnAB. Nat Rev Microbiol 11, 9–13 (2013). https://doi.org/10.1038/nrmicro2917
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2917
This article is cited by
-
Complete Genome Sequencing Analysis of Deinococcus wulumuqiensis R12, an Extremely Radiation-Resistant Strain
Current Microbiology (2022)
-
Crystal structure of the nuclease and capping domain of SbcD from Staphylococcus aureus
Journal of Microbiology (2021)
-
DNA targeting and interference by a bacterial Argonaute nuclease
Nature (2020)
-
Microbial ageing and longevity
Nature Reviews Microbiology (2019)
-
Modeling DNA Unwinding by AddAB Helicase–Nuclease and Modulation by Chi Sequences: Comparison with AdnAB and RecBCD
Cellular and Molecular Bioengineering (2019)