Key Points
-
To evaluate current progress in GPCR structure prediction and ligand docking, a community-wide prediction assessment — GPCR Dock 2008 — in coordination with the publication of the human adenosine A2A receptor structure in October 2008 and public release of the 3-dimensional coordinates.
-
Twenty-nine groups submitted 206 structural models before the release of the experimental structure. The structures were evaluated for the accuracy of the ligand binding mode and the overall receptor model compared with the crystal structure.
-
The majority of the submitted models predicted the overall topology, but did not predict the ligand position and the binding interactions very accurately.
-
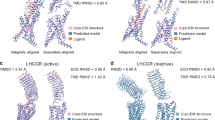
The best model overall (submitted by S. Costanzi) has a ligand RMSD of 2.8 Å RMSD and 34 of 75 correct contacts.
-
Accurate modelling of the structurally divergent regions (such as the extracellular loops), of disulphide bond formation affecting helix residue registry and of the helical shifts in the TM region seem to be crucial for accurately predicting the key ligand interactions in GPCRs, and this area is perhaps the most in need of technological development.
Abstract
Recent breakthroughs in the determination of the crystal structures of G protein-coupled receptors (GPCRs) have provided new opportunities for structure-based drug design strategies targeting this protein family. With the aim of evaluating the current status of GPCR structure prediction and ligand docking, a community-wide, blind prediction assessment — GPCR Dock 2008 — was conducted in coordination with the publication of the crystal structure of the human adenosine A2A receptor bound to the ligand ZM241385. Twenty-nine groups submitted 206 structural models before the release of the experimental structure, which were evaluated for the accuracy of the ligand binding mode and the overall receptor model compared with the crystal structure. This analysis highlights important aspects for success and future development, such as accurate modelling of structurally divergent regions and use of additional biochemical insight such as disulphide bridges in the extracellular loops.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jorgensen, W. L. The many roles of computation in drug discovery. Science 303, 1813–1818 (2004).
Richon, A. B. Current status and future direction of the molecular modeling industry. Drug Discov. Today 13, 665–669 (2008).
Kitchen, D. B., Decornez, H., Furr, J. R. & Bajorath, J. Docking and scoring in virtual screening for drug discovery: methods and applications. Nature Rev. Drug Discov. 3, 935–949 (2004).
Drews, J. Drug discovery: a historical perspective. Science 287, 1960–1964 (2000).
Klabunde, T. & Hessler, G. Drug design strategies for targeting G-protein-coupled receptors. Chembiochem. 3, 928–944 (2002).
Becker, O. M. et al. G protein-coupled receptors: in silico drug discovery in 3D. Proc. Natl Acad. Sci. USA 101, 11304–11309 (2004).
Ballesteros, J. & Palczewski, K. G protein-coupled receptor drug discovery: implications from the crystal structure of rhodopsin. Curr. Opin. Drug Discov. Devel. 4, 561–574 (2001).
Bu, L., Michino, M., Wolf, R. M. & Brooks, C. L. III. Improved model building and assessment of the Calcium-sensing receptor transmembrane domain. Proteins 71, 215–226 (2008).
Henin, J. et al. Probing a model of a GPCR/ligand complex in an explicit membrane environment: the human cholecystokinin-1 receptor. Biophys. J. 90, 1232–1240 (2006).
Fowler, C. B., Pogozheva, I. D., LeVine, H., 3rd & Mosberg, H. I. Refinement of a homology model of the mu-opioid receptor using distance constraints from intrinsic and engineered zinc-binding sites. Biochemistry 43, 8700–8710 (2004).
Evers, A. & Klabunde, T. Structure-based drug discovery using GPCR homology modeling: successful virtual screening for antagonists of the alpha1A adrenergic receptor. J. Med. Chem. 48, 1088–1097 (2005).
Manivet, P. et al. The serotonin binding site of human and murine 5-HT2B receptors: molecular modeling and site-directed mutagenesis. J. Biol. Chem. 277, 17170–17178 (2002).
Archer, E., Maigret, B., Escrieut, C., Pradayrol, L. & Fourmy, D. Rhodopsin crystal: new template yielding realistic models of G-protein-coupled receptors? Trends Pharmacol. Sci. 24, 36–40 (2003).
Gershengorn, M. C. & Osman, R. Minireview: Insights into G protein-coupled receptor function using molecular models. Endocrinology 142, 2–10 (2001).
Ballesteros, J. A., Shi, L. & Javitch, J. A. Structural mimicry in G protein-coupled receptors: implications of the high-resolution structure of rhodopsin for structure-function analysis of rhodopsin-like receptors. Mol. Pharmacol. 60, 1–19 (2001).
Kobilka, B. K. & Deupi, X. Conformational complexity of G-protein-coupled receptors. Trends Pharmacol. Sci. 28, 397–406 (2007).
Bhattacharya, S., Hall, S. E., Li, H. & Vaidehi, N. Ligand-stabilized conformational states of human beta(2) adrenergic receptor: insight into G-protein-coupled receptor activation. Biophys. J. 94, 2027–2042 (2008).
Kenakin, T. Efficacy at G-protein-coupled receptors. Nature Rev. Drug Discov. 1, 103–110 (2002).
Jaakola, V. P. et al. The 2.6 angstrom crystal structure of a human A2A adenosine receptor bound to an antagonist. Science 322, 1211–1217 (2008). The human adenosine A 2A receptor crystal structure served as the experimental template for comparison for this modelling and docking assessment. This is the second human GPCR structure to be experimentally determined.
Jacobson, K. A. & Gao, Z. G. Adenosine receptors as therapeutic targets. Nature Rev. Drug Discov. 5, 247–264 (2006).
Moult, J. et al. Critical assessment of methods of protein structure prediction-Round VII. Proteins 69 (Suppl. 8), 3–9 (2007). The very successful CASP (Critical Assessment of Protein Structure) project started in 1994 and served as the model to conduct the reported GPCR Dock 2008 modelling and docking assessment.
Lensink, M. F., Mendez, R. & Wodak, S. J. Docking and scoring protein complexes: CAPRI 3rd Edition. Proteins 69, 704–718 (2007).
Kopp, J., Bordoli, L., Battey, J. N., Kiefer, F. & Schwede, T. Assessment of CASP7 predictions for template-based modeling targets. Proteins 69 (Suppl. 8), 38–56 (2007).
Kobilka, B. & Schertler, G. F. New G-protein-coupled receptor crystal structures: insights and limitations. Trends Pharmacol. Sci. 29, 79–83 (2008).
Jacobson, M. P. et al. A hierarchical approach to all-atom protein loop prediction. Proteins 55, 351–367 (2004).
Rohl, C. A., Strauss, C. E., Chivian, D. & Baker, D. Modeling structurally variable regions in homologous proteins with rosetta. Proteins 55, 656–677 (2004).
Friesner, R. A. et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J. Med. Chem. 47, 1739–1749 (2004).
Totrov, M. & Abagyan, R. Flexible protein-ligand docking by global energy optimization in internal coordinates. Proteins 29 (Suppl. 1), 215–220 (1997).
Verdonk, M. L., Cole, J. C., Hartshorn, M. J., Murray, C. W. & Taylor, R. D. Improved protein-ligand docking using GOLD. Proteins 52, 609–623 (2003).
Morris, G. et al. Automated docking using a lamarkian genetic algorithm and empirical binding free energy function. J. Comput. Chem. 19, 1639–1662 (1998).
Mirzadegan, T., Benko, G., Filipek, S. & Palczewski, K. Sequence analyses of G-protein-coupled receptors: similarities to rhodopsin. Biochemistry 42, 2759–2767 (2003).
Baker, D. & Sali, A. Protein structure prediction and structural genomics. Science 294, 93–96 (2001). The use of protein models and docking is dependent on how such data will be used. In this paper, Baker and Sali provide an excellent presentation of where models are useful, in particular as hypothesis generators with the application being dependent on the resolution of the structure.
Fredriksson, R., Lagerstrom, M. C., Lundin, L. G. & Schioth, H. B. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 63, 1256–1272 (2003).
Marti-Renom, M. A. et al. Comparative protein structure modeling of genes and genomes. Annu. Rev. Biophys. Biomol. Struct. 29, 291–325 (2000).
Read, R. J. & Chavali, G. Assessment of CASP7 predictions in the high accuracy template-based modeling category. Proteins 69 (Suppl. 8), 27–37 (2007).
Kim, J. et al. Glutamate residues in the second extracellular loop of the human A2a adenosine receptor are required for ligand recognition. Mol. Pharmacol. 49, 683–691 (1996).
Barth, P., Schonbrun, J. & Baker, D. Toward high-resolution prediction and design of transmembrane helical protein structures. Proc. Natl Acad. Sci. USA 104, 15682–15687 (2007).
Acknowledgements
We thank M. Hanson, V.-P. Jaakola, C. Roth and V. Cherezov for help with the analysis and comments on the manuscript, and K. Kadyshevskaya and V. Cherezov for figure preparation. We are grateful to the Goddard group for providing the script to calculate the binding site contact RMSD. We thank A. Walker for data tracking and assistance with the manuscript and J. Kunken for IT help during the assessment. This work was supported in part by the Protein Structure Initiative grant U54 GM074961 (ATCG3D), the NIH Roadmap grant P50 GM073197 (JCIMPT), and the Multiscale Modeling Tools for Structural Biology NCRR via grant P41 RR012255.
Author information
Authors and Affiliations
Consortia
Corresponding author
Supplementary information
Supplementary information S1 (box)
Computational methods used by the GPCR Dock 2008 participants (PDF 3365 kb)
Glossary
- Rhodopsin and bacteriorhodopsin
-
These two light-activated membrane proteins have a seven transmembrane alpha-helical bundle architecture that is similar to the general structure of the larger GPCR family.
- Molecular dynamics simulation
-
This molecular modelling approach uses numerical integration to solve the equations of motion based on the forces arising from interatomic interactions. The dynamic behaviour of atoms in a macromolecular system, such as that in a membrane protein, can be understood by running a molecular dynamics (MD) simulation. MD simulation can also be used to refine structural models of proteins and protein–ligand complexes.
- Ligand docking
-
A molecular modelling approach that predicts the ligand binding mode within a targeted binding site. In this approach, the known or predicted three-dimensional structure of a protein is probed using computationally generated energy landscapes to identify the most favourable binding pose for the ligand.
- RMSD (root mean square deviation)
-
RMSD is used as a quantitative measure of the similarity between two superimposed atomic coordinates. RMSD values (units of Å) can be calculated for any type and subset of atoms; for example, Cα atoms of proteins (Cα RMSD) for all residues, for residues in the transmembrane helices or the loops; heavy atoms of small-molecule ligands (ligand RMSD).
- Z-score
-
A standard dimensionless score that normalizes a value with respect to the sample mean and standard deviation.
- Cα atoms
-
The chiral carbon atoms to which the primary amine, the carboxylic group and the side chain are attached to in an amino acid. Comparison of three-dimensional structures of proteins is sometimes carried out by superimposing the Cα atoms of proteins as this provides a simple estimate of the similarity of their skeleton or backbone structure.
- B-factor
-
A descriptor that reflects the fluctuation of atomic position from an atom's average position and provides important insight into a protein's potential dynamic behaviour.
- Hydrogen bond
-
Attractive interaction between one electronegative atom and a hydrogen covalently bonded to another electronegative atom such as nitrogen or oxygen.
- Aromatic stacking
-
Attractive interactions between the aromatic rings of amino acids. Overlapping of p-orbitals of π-conjugated systems result in the rings arranging themselves in preferred orientations.
Rights and permissions
About this article
Cite this article
Michino, M., Abola, E., GPCR Dock 2008 participants. et al. Community-wide assessment of GPCR structure modelling and ligand docking: GPCR Dock 2008. Nat Rev Drug Discov 8, 455–463 (2009). https://doi.org/10.1038/nrd2877
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd2877
This article is cited by
-
Benchmarking the performance of MM/PBSA in virtual screening enrichment using the GPCR-Bench dataset
Journal of Computer-Aided Molecular Design (2020)
-
Dual binding mode of “bitter sugars” to their human bitter taste receptor target
Scientific Reports (2019)
-
A benchmark study of loop modeling methods applied to G protein-coupled receptors
Journal of Computer-Aided Molecular Design (2019)
-
Evaluating the performance of MM/PBSA for binding affinity prediction using class A GPCR crystal structures
Journal of Computer-Aided Molecular Design (2019)
-
Structure-Activity Investigations and Optimisations of Non-metabolite Agonists for the Succinate Receptor 1
Scientific Reports (2018)