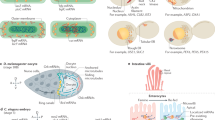
Abstract
This protocol describes a nonisotopic method for high-resolution investigation of the kinetics of RNA within the cell. This involves the incorporation of bromouridine-5′-triphosphate into RNA of living cells by lipofection followed by immunocytological detection of BrRNAs. The use of the same antibody identified either with fluorescence or with gold particles revealed the three-dimensional organization of sites containing labeled RNAs or their precise localization by using confocal and ultrastructural microscopy, respectively. Comparison of three-dimensional reconstruction obtained from the series of optical sections and ultrathin sections was extremely fruitful to describe topological and spatial dynamics of RNAs from their synthesis site inside the nucleus to the cytoplasm. Combined with immunolocalization of proteins involved in different nuclear activities and with highly resolved three-dimensional visualizations of the labelings, this method should also provide a significant contribution to our understanding of the functional, volumic organization of the cell nucleus. The entire protocol can be completed in ∼10 d.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Fakan, S. High resolution autoradiography studies on chromatin functions. In The Cell Nucleus Vol. 5 (ed. Busch, H.) 3–53 (Academic press, New York, USA, 1978).
Jackson, D.A., Hassan, A.B., Errington, R.J. & Cook, P.R. Visualization of focal sites of transcription within human nuclei. EMBO J. 12, 1059–1065 (1993).
Wansink, D.G. et al. Fluorescent labeling of nascent RNA reveals transcription by RNA polymerase II in domains scattered throughout the nucleus. J. Cell Biol. 122, 283–293 (1993).
Dundr, M. & Raska, I. Nonisotopic ultrastructural mapping of transcription sites within the nucleolus. Exp. Cell Res. 208, 275–281 (1993).
Hozák, P., Cook, P.R., Schofer, C., Mosgoller, W. & Wachtler, F. Site of transcription of ribosomal RNA and intranucleolar structure in HeLa cells. J. Cell Sci. 107, 639–648 (1994).
Masson, C. et al. Conditions favoring RNA polymerase I transcription in permeabilized cells. Exp. Cell Res. 226, 114–125 (1996).
Koberna, K. et al. Nuclear organization studied with the help of a hypotonic shift: its use permits hydrophilic molecules to enter into living cells. Chromosoma 108, 325–335 (1999).
Koberna, K. et al. Ribosomal genes in focus: new transcripts label the dense fibrillar components and form clusters indicative of 'Christmas trees' in situ . J. Cell Biol. 157, 743–748 (2002).
Cmarko, D. et al. Ultrastructural analysis of nucleolar transcription in cells microinjected with 5-bromo-UTP. Histochem. Cell Biol. 113, 181–187 (2000).
Fay, F.S., Taneja, K.L., Shenoy, S., Lifshitz, L. & Singer, R.H. Quantitative digital analysis of diffuse and concentrated nuclear distributions of nascent transcripts, SC35 and poly(A). Exp. Cell Res. 231, 27–37 (1997).
Iborra, F.J., Jackson, D.A. & Cook, P.R. The path of transcripts from extra-nucleolar synthetic sites to nuclear pores: transcripts in transit are concentrated in discrete structures containing SR proteins. J. Cell Sci. 111, 2269–2282 (1998).
Melcák, I., Risueno, M.C. & Raska, I. Ultrastructural nonisotopic mapping of nucleolar transcription sites in onion protoplasts. J. Struct. Biol. 116, 253–263 (1996).
Thiry, M., Cheutin, T., O'Donohue, M.F., Kaplan, H. & Ploton, D. Dynamics and three-dimensional localization of ribosomal RNA within the nucleolus. RNA 6, 1750–1761 (2000).
Haukenes, G., Szilvay, A.M., Brokstad, K.A., Kanestrom, A. & Kalland, K.H. Labeling of RNA transcripts of eukaryotic cells in culture with BrUTP using a liposome transfection reagent (DOTAP). BioTechniques 21, 308–312 (1997).
Kanestrom, A., Andresen, V., Szilvay, A.M., Kalland, K.H. & Haukenes, G. Histographic recording of human immunodeficiency virus type 1 (HIV-1) regulatory protein Rev and nuclear factors. Arch. Virol. 143, 279–294 (1998).
Malínskỳ, J., Karel Koberna, K., Bednár, J. & Stulík, J. Raska I. Searching for active ribosomal genes in situ: light microscopy in light of the electron beam. J. Struct. Biol. 40, 227–231 (2002).
Raska, I. Oldies but goldies: searching for Christmas trees within the nucleolar architecture. Trends Cell Biol. 10, 517–525 (2003).
Acknowledgements
We thank Miss F. Skivée for their skillful technical assistance. This work received financial support from the 'Fonds National de la Recherche Scientifique Médicale' (grant no. 3.4540.06), ARC (contract no. 4497) and Ligue contre le Cancer (Departements de l'Aube, de la Marne et des Ardennes). F.L. and N.T. are PhD grant holders of the FNRS.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
41596_2008_BFnprot2008198_MOESM352_ESM.mov
Supplementary Video 1: Volume rendering of Br-RNAs after a 15 min pulse within a multinucleated cell (without α-amanitin) showing a strong nucleolar, nucleoplasmic and cytoplasmic labelling. (MOV 2602 kb)
41596_2008_BFnprot2008198_MOESM353_ESM.mov
Supplementary Video 2: Volume rendering of Br-RNAs after a 15 min pulse within two α-amanitin-treated cells showing a strong nucleolar and cytoplasmic labelling. (MOV 1869 kb)
41596_2008_BFnprot2008198_MOESM354_ESM.mov
Supplementary Video 3: Surface rendering of Br-RNAs and RNA polymerase 1 after a 5 min pulse within a α-amanitin treated cell. Br-RNAs (solid green), RNA polymerase 1 (solid red) are shown alone or simultaneously on left, middle and right panels respectively. The two labellings appear as totally overlapping individual full beads 0.5 to 1 µm in diameter, organized as several necklaces. (MOV 2194 kb)
41596_2008_BFnprot2008198_MOESM355_ESM.mov
Supplementary Video 4: Surface rendering of Br-RNAs and RNA polymerase 1 after a 5 min pulse within a α-amanitin treated cell (same cell as on movie 3). Br-RNAs (transparent green) and RNA polymerase 1 (solid red) are shown simultaneously (left panel). Voxels with co-localized high levels of green and red signals were computed and shown in solid blue either alone (middle panel) or simultaneously with Br-RNAs (transparent green) (right panel). Note that co-localized voxels represent 95% of voxels containing high levels of BrRNA labelling and that they are located in its central part. (MOV 2709 kb)
41596_2008_BFnprot2008198_MOESM356_ESM.mov
Supplementary Video 5: Surface rendering of Br-RNAs and RNA polymerase 1 after a 15 min pulse within a α-amanitin treated cell. Br-RNAs (solid green), RNA polymerase 1 (solid red) are shown alone or simultaneously on left, middle and right panels respectively. Br-RNAs labelling is disposed as large hollow beads partly overlapping RNA polymerase 1 labelling. The latter is disposed as individual full beads 0.5 to 1 µm in diameter, organized as several necklaces. (MOV 2590 kb)
41596_2008_BFnprot2008198_MOESM357_ESM.mov
Supplementary Video 6: Surface rendering of Br-RNAs and RNA polymerase 1 after a 15 min pulse within a α-amanitin treated cell (same cell as on movie 5). Br-RNAs (transparent green) and RNA polymerase 1 (solid red) are shown simultaneously (left panel). Voxels with co-localized high levels of green and red signals were computed and shown in solid blue either alone (middle panel) or simultaneously with Br-RNAs (transparent green) (right panel). Note that co-localized voxels represent 69% of voxels containing high levels of BrRNA labelling and that they are located in its central part. (MOV 2827 kb)
41596_2008_BFnprot2008198_MOESM358_ESM.mov
Supplementary Video 7: Surface rendering of Br-RNAs and RNA polymerase 1 after a 30 min pulse within a α-amanitin treated cell. Br-RNAs (solid green), RNA polymerase 1 (solid red) are shown alone or simultaneously on left, middle and right panels respectively. Br-RNAs labelling constitutes large sponge-like structures with many holes. Some of the latter contain RNA polymerase 1 labelling which is disposed as individual full beads 0.5 to 1 µm in diameter and organized as several necklaces. (MOV 2785 kb)
41596_2008_BFnprot2008198_MOESM359_ESM.mov
Supplementary Video 8: Surface rendering of Br-RNAs and RNA polymerase 1 after a 30 min pulse within a α-amanitin treated cell (same cell as on movie 7). Br-RNAs (transparent green) and RNA polymerase 1 (solid red) are shown simultaneously (left panel). Voxels with co-localized high levels of green and red signals were computed and shown in solid blue either alone (middle panel) or simultaneously with Br-RNAs (transparent green) (right panel). Note that co-localized voxels represent only 15% of voxels containing high levels of BrRNA labelling. (MOV 2593 kb)
Rights and permissions
About this article
Cite this article
Thiry, M., Lamaye, F., Thelen, N. et al. A protocol for studying the kinetics of RNA within cultured cells: application to ribosomal RNA. Nat Protoc 3, 1997–2004 (2008). https://doi.org/10.1038/nprot.2008.198
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2008.198
This article is cited by
-
RNA content in the nucleolus alters p53 acetylation via MYBBP1A
The EMBO Journal (2011)
-
Localization of Nopp140 within mammalian cells during interphase and mitosis
Histochemistry and Cell Biology (2009)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.