Abstract
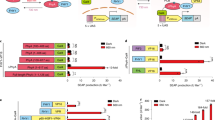
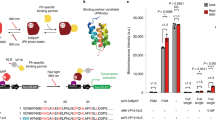
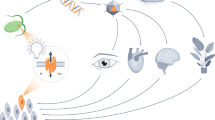
Light-mediated control of protein–protein interactions to regulate cellular pathways is an important application of optogenetics. Here, we report an optogenetic system based on the reversible light-induced binding between the bacterial phytochrome BphP1 and its natural partner PpsR2 from Rhodopseudomonas palustris bacteria. We extensively characterized the BphP1–PpsR2 interaction both in vitro and in mammalian cells and then used this interaction to translocate target proteins to specific cellular compartments, such as the plasma membrane and the nucleus. We showed light-inducible control of cell morphology that resulted in a substantial increase of the cell area. We demonstrated light-dependent gene expression with 40-fold contrast in cultured cells, 32-fold in subcutaneous mouse tissue, and 5.7-fold in deep tissues in mice. Characteristics of the BphP1–PpsR2 optogenetic system include its sensitivity to 740- to 780-nm near-infrared light, its ability to utilize an endogenous biliverdin chromophore in eukaryotes (including mammals), and its spectral compatibility with blue-light-driven optogenetic systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Shcherbakova, D.M., Shemetov, A.A., Kaberniuk, A.A. & Verkhusha, V.V. Natural photoreceptors as a source of fluorescent proteins, biosensors, and optogenetic tools. Annu. Rev. Biochem. 84, 519–550 (2015).
Motta-Mena, L.B. et al. An optogenetic gene expression system with rapid activation and deactivation kinetics. Nat. Chem. Biol. 10, 196–202 (2014).
Kawano, F., Suzuki, H., Furuya, A. & Sato, M. Engineered pairs of distinct photoswitches for optogenetic control of cellular proteins. Nat. Commun. 6, 6256 (2015).
Stierl, M. et al. Light modulation of cellular cAMP by a small bacterial photoactivated adenylyl cyclase, bPAC, of the soil bacterium Beggiatoa. J. Biol. Chem. 286, 1181–1188 (2011).
Taslimi, A. et al. An optimized optogenetic clustering tool for probing protein interaction and function. Nat. Commun. 5, 4925 (2014).
Lee, S. et al. Reversible protein inactivation by optogenetic trapping in cells. Nat. Methods 11, 633–636 (2014).
Ni, M., Tepperman, J.M. & Quail, P.H. Binding of phytochrome B to its nuclear signalling partner PIF3 is reversibly induced by light. Nature 400, 781–784 (1999).
Levskaya, A., Weiner, O.D., Lim, W.A. & Voigt, C.A. Spatiotemporal control of cell signalling using a light-switchable protein interaction. Nature 461, 997–1001 (2009).
Gomez, E.J., Gerhardt, K., Judd, J., Tabor, J.J. & Suh, J. Light-activated nuclear translocation of adeno-associated virus nanoparticles using phytochrome B for enhanced, tunable, and spatially programmable gene delivery. ACS Nano. 10, 225–237 (2016).
Weissleder, R. & Ntziachristos, V. Shedding light onto live molecular targets. Nat. Med. 9, 123–128 (2003).
Ulijasz, A.T. & Vierstra, R.D. Phytochrome structure and photochemistry: recent advances toward a complete molecular picture. Curr. Opin. Plant Biol. 14, 498–506 (2011).
Piatkevich, K.D., Subach, F.V. & Verkhusha, V.V. Engineering of bacterial phytochromes for near-infrared imaging, sensing, and light-control in mammals. Chem. Soc. Rev. 42, 3441–3452 (2013).
Müller, K. et al. A red/far-red light-responsive bi-stable toggle switch to control gene expression in mammalian cells. Nucleic Acids Res. 41, e77 (2013).
Tran, M.T. et al. In vivo image analysis using iRFP transgenic mice. Exp. Anim. 63, 311–319 (2014).
Shcherbakova, D.M., Baloban, M. & Verkhusha, V.V. Near-infrared fluorescent proteins engineered from bacterial phytochromes. Curr. Opin. Chem. Biol. 27, 52–63 (2015).
Shcherbakova, D.M. & Verkhusha, V.V. Near-infrared fluorescent proteins for multicolor in vivo imaging. Nat. Methods 10, 751–754 (2013).
Shcherbakova, D.M. et al. Molecular basis of spectral diversity in near-infrared phytochrome-based fluorescent proteins. Chem. Biol. 22, 1540–1551 (2015).
Piatkevich, K.D., Subach, F.V. & Verkhusha, V.V. Far-red light photoactivatable near-infrared fluorescent proteins engineered from a bacterial phytochrome. Nat. Commun. 4, 2153 (2013).
Filonov, G.S. & Verkhusha, V.V. A near-infrared BiFC reporter for in vivo imaging of protein-protein interactions. Chem. Biol. 20, 1078–1086 (2013).
Auldridge, M.E. & Forest, K.T. Bacterial phytochromes: more than meets the light. Crit. Rev. Biochem. Mol. Biol. 46, 67–88 (2011).
Ryu, M.H. & Gomelsky, M. Near-infrared light responsive synthetic c-di-GMP module for optogenetic applications. ACS Synth. Biol. 3, 802–810 (2014).
Ryu, M.H. et al. Engineering adenylate cyclases regulated by near-infrared window light. Proc. Natl. Acad. Sci. USA 111, 10167–10172 (2014).
Gasser, C. et al. Engineering of a red-light-activated human cAMP/cGMP-specific phosphodiesterase. Proc. Natl. Acad. Sci. USA 111, 8803–8808 (2014).
Wagner, J.R., Zhang, J., Brunzelle, J.S., Vierstra, R.D. & Forest, K.T. High resolution structure of Deinococcus bacteriophytochrome yields new insights into phytochrome architecture and evolution. J. Biol. Chem. 282, 12298–12309 (2007).
Rockwell, N.C. & Lagarias, J.C. A brief history of phytochromes. ChemPhysChem 11, 1172–1180 (2010).
Rottwinkel, G., Oberpichler, I. & Lamparter, T. Bathy phytochromes in rhizobial soil bacteria. J. Bacteriol. 192, 5124–5133 (2010).
Kojadinovic, M. et al. Dual role for a bacteriophytochrome in the bioenergetic control of Rhodopseudomonas palustris: enhancement of photosystem synthesis and limitation of respiration. Biochim. Biophys. Acta 1777, 163–172 (2008).
Bellini, D. & Papiz, M.Z. Structure of a bacteriophytochrome and light-stimulated protomer swapping with a gene repressor. Structure 20, 1436–1446 (2012).
Hussain, N.K. et al. Endocytic protein intersectin-l regulates actin assembly via Cdc42 and N-WASP. Nat. Cell Biol. 3, 927–932 (2001).
Hall, A. Rho GTPases and the actin cytoskeleton. Science 279, 509–514 (1998).
Nobes, C.D. & Hall, A. Rho, rac, and cdc42 GTPases regulate the assembly of multimolecular focal complexes associated with actin stress fibers, lamellipodia, and filopodia. Cell 81, 53–62 (1995).
Orth, P., Schnappinger, D., Hillen, W., Saenger, W. & Hinrichs, W. Structural basis of gene regulation by the tetracycline inducible Tet repressor-operator system. Nat. Struct. Biol. 7, 215–219 (2000).
Wang, X., Chen, X. & Yang, Y. Spatiotemporal control of gene expression by a light-switchable transgene system. Nat. Methods 9, 266–269 (2012).
Chen, X., Li, T., Wang, X. & Yang, Y. A light-switchable bidirectional expression module allowing simultaneous regulation of multiple genes. Biochem. Biophys. Res. Commun. 465, 769–776 (2015).
Schönig, K., Bujard, H. & Gossen, M. The power of reversibility regulating gene activities via tetracycline-controlled transcription. Methods Enzymol. 477, 429–453 (2010).
Albanese, C., Hulit, J., Sakamaki, T. & Pestell, R.G. Recent advances in inducible expression in transgenic mice. Semin. Cell Dev. Biol. 13, 129–141 (2002).
Toettcher, J.E., Weiner, O.D. & Lim, W.A. Using optogenetics to interrogate the dynamic control of signal transmission by the Ras/Erk module. Cell 155, 1422–1434 (2013).
Müller, K. et al. Synthesis of phycocyanobilin in mammalian cells. Chem. Commun. (Camb.) 49, 8970–8972 (2013).
International Electrotechnical Commission. Safety of Laser Products–Part 1: Equipment Classification and Requirements 3rd edn. (International Electrotechnical Commission, 2014).
Lam, A.J. et al. Improving FRET dynamic range with bright green and red fluorescent proteins. Nat. Methods 9, 1005–1012 (2012).
Cui, Z., Geurts, A.M., Liu, G., Kaufman, C.D. & Hackett, P.B. Structure-function analysis of the inverted terminal repeats of the sleeping beauty transposon. J. Mol. Biol. 318, 1221–1235 (2002).
Koh, E.Y. et al. An internal ribosome entry site (IRES) mutant library for tuning expression level of multiple genes in mammalian cells. PLoS ONE 8, e82100 (2013).
Liu, F., Song, Y. & Liu, D. Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA. Gene Ther. 6, 1258–1266 (1999).
Acknowledgements
We thank E. Giraud (Institute for Research and Development, Marseille), M. Papiz (Liverpool University), T. Beatty (University of British Columbia), W. Weber (University of Freiburg), P. Hackett (University of Minnesota), Z. Izsvak (Max Delbrück Center for Molecular Medicine), Y. Yang (East China University of Science and Technology), and S. Masuda (Tokyo Institute of Technology) for plasmids and D. Shcherbakova, K. Chernov, and T. Redchuk for useful suggestions. This work was sponsored by National Institutes of Health grants GM073913, GM108579, and CA164468 to V.V.V.
Author information
Authors and Affiliations
Contributions
A.A.K. and A.A.S. characterized the proteins in vitro, in mammalian cell culture, and in vivo. V.V.V. planned and directed the project and, together with A.A.K. and A.A.S., designed the experiments, analyzed the data, and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Tables 1 and 2, and Supplementary Note (PDF 2453 kb)
Rights and permissions
About this article
Cite this article
Kaberniuk, A., Shemetov, A. & Verkhusha, V. A bacterial phytochrome-based optogenetic system controllable with near-infrared light. Nat Methods 13, 591–597 (2016). https://doi.org/10.1038/nmeth.3864
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3864
This article is cited by
-
Controlling cellular activities with light
Nature Methods (2023)
-
A small and highly sensitive red/far-red optogenetic switch for applications in mammals
Nature Biotechnology (2022)
-
A red light–responsive photoswitch for deep tissue optogenetics
Nature Biotechnology (2022)
-
Optogenetic manipulation and photoacoustic imaging using a near-infrared transgenic mouse model
Nature Communications (2022)
-
Synthetic protein condensates for cellular and metabolic engineering
Nature Chemical Biology (2022)