Abstract
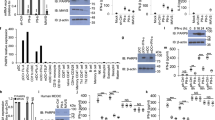
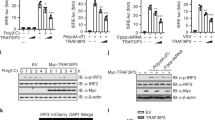
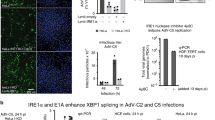
Viral infection triggers innate immune sensors to produce type I interferon. However, infection of T cells and macrophages with human immunodeficiency virus (HIV) does not trip those alarms. How HIV avoids activating nucleic acid sensors is unknown. Here we found that the cytosolic exonuclease TREX1 suppressed interferon triggered by HIV. In Trex1−/− mouse cells and human CD4+ T cells and macrophages in which TREX1 was inhibited by RNA-mediated interference, cytosolic HIV DNA accumulated and HIV infection induced type I interferon that inhibited HIV replication and spreading. TREX1 bound to cytosolic HIV DNA and digested excess HIV DNA that would otherwise activate interferon expression via a pathway dependent on the kinase TBK1, the adaptor STING and the transcription factor IRF3. HIV-stimulated interferon production in cells deficient in TREX1 did not involve known nucleic acid sensors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bowerman, B., Brown, N., Bishop, K. & Varmus, H. A nucleoprotein complex mediates the integration of retroviral DNA. Genes Dev. 3, 469–478 (1989).
Yan, N., Cherepanov, P., Daigle, J.E., Engelman, A. & Lieberman, J. The SET complex acts as a barrier to autointegration of HIV-1. PLoS Pathog. 5, e1000327 (2009).
Lee-Kirsch, M.A. et al. A mutation in TREX1 that impairs susceptibility to granzyme A-mediated cell death underlies familial chilblain lupus. J. Mol. Med. 85, 531–537 (2007).
Lee-Kirsch, M.A. et al. Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 are associated with systemic lupus erythematosus. Nat. Genet. 39, 1065–1067 (2007).
Crow, Y. et al. Mutations in the gene encoding the 3′-5′ DNA exonuclease TREX1 cause Aicardi-Goutieres syndrome at the AGS1 locus. Nat. Genet. 38, 917–920 (2006).
Stetson, D.B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Stetson, D.B., Ko, J.S., Heidmann, T. & Medzhitov, R. Trex1 prevents cell-intrinsic initiation of autoimmunity. Cell 134, 587–598 (2008).
Yang, Y., Lindahl, T. & Barnes, D. Trex1 exonuclease degrades ssDNA to prevent chronic checkpoint activation and autoimmune disease. Cell 131, 873–886 (2007).
Shun, M. et al. LEDGF/p75 functions downstream from preintegration complex formation to effect gene-specific HIV-1 integration. Genes Dev. 21, 1767–1778 (2007).
Honda, K. et al. IRF-7 is the master regulator of type-I interferon-dependent immune responses. Nature 434, 772–777 (2005).
Kawai, T. et al. Interferon-α induction through Toll-like receptors involves a direct interaction of IRF7 with MyD88 and TRAF6. Nat. Immunol. 5, 1061–1068 (2004).
Agy, M.B., Acker, R.L., Sherbert, C.H. & Katze, M.G. Interferon treatment inhibits virus replication in HIV-1- and SIV-infected CD4+ T-cell lines by distinct mechanisms: evidence for decreased stability and aberrant processing of HIV-1 proteins. Virology 214, 379–386 (1995).
Coccia, E.M., Krust, B. & Hovanessian, A.G. Specific inhibition of viral protein synthesis in HIV-infected cells in response to interferon treatment. J. Biol. Chem. 269, 23087–23094 (1994).
Shirazi, Y. & Pitha, P.M. Alpha interferon inhibits early stages of the human immunodeficiency virus type 1 replication cycle. J. Virol. 66, 1321–1328 (1992).
Baca-Regen, L., Heinzinger, N., Stevenson, M. & Gendelman, H.E. Alpha interferon-induced antiretroviral activities: restriction of viral nucleic acid synthesis and progeny virion production in human immunodeficiency virus type 1-infected monocytes. J. Virol. 68, 7559–7565 (1994).
Mazur, D. & Perrino, F. Identification and expression of the TREX1 and TREX2 cDNA sequences encoding mammalian 3′→5′ exonucleases. J. Biol. Chem. 274, 19655–19660 (1999).
Lehtinen, D.A., Harvey, S., Mulcahy, M.J., Hollis, T. & Perrino, F.W. The TREX1 double-stranded DNA degradation activity is defective in dominant mutations associated with autoimmune disease. J. Biol. Chem. 283, 31649–31656 (2008).
Hornung, V. & Latz, E. Intracellular DNA recognition. Nat. Rev. Immunol. 10, 123–130 (2010).
Schroder, K., Muruve, D.A. & Tschopp, J. Innate immunity: cytoplasmic DNA sensing by the AIM2 inflammasome. Curr. Biol. 19, R262–R265 (2009).
Yang, P. et al. The cytosolic nucleic acid sensor LRRFIP1 mediates the production of type I interferon via a β-catenin-dependent pathway. Nat. Immunol. 11, 487–494 (2010).
Yanai, H. et al. HMGB proteins function as universal sentinels for nucleic-acid-mediated innate immune responses. Nature 462, 99–103 (2009).
Chiu, Y.-H., Macmillan, J.B. & Chen, Z.J. RNA polymerase III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell 138, 576–591 (2009).
Ablasser, A. et al. RIG-I-dependent sensing of poly(dA:dT) through the induction of an RNA polymerase III-transcribed RNA intermediate. Nat. Immunol. 10, 1065–1072 (2009).
Seth, R.B., Sun, L., Ea, C.-K. & Chen, Z.J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-kappaB and IRF 3. Cell 122, 669–682 (2005).
Kawai, T. et al. IPS-1, an adaptor triggering RIG-I- and Mda5-mediated type I interferon induction. Nat. Immunol. 6, 981–988 (2005).
Xu, L.-G. VISA is an adapter protein required for virus-triggered IFN-β signaling. Mol. Cell 19, 727–740 (2005).
Meylan, E. et al. Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus. Nature 437, 1167–1172 (2005).
Ishikawa, H. & Barber, G. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 647–678 (2008).
Zhong, B. et al. The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29, 538–550 (2008).
Sun, W. et al. ERIS, an endoplasmic reticulum IFN stimulator, activates innate immune signaling through dimerization. Proc. Natl. Acad. Sci. USA 106, 8653–8658 (2009).
Ishikawa, H., Ma, Z. & Barber, G.N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Fan, Z., Beresford, P., Zhang, D. & Lieberman, J. HMG2 interacts with the nucleosome assembly protein SET and is a target of the cytotoxic T-lymphocyte protease granzyme A. Mol. Cell. Biol. 22, 2810–2820 (2002).
Lehming, N., Le Saux, A., Schüller, J. & Ptashne, M. Chromatin components as part of a putative transcriptional repressing complex. Proc. Natl. Acad. Sci. USA 95, 7322–7326 (1998).
Gabellini, D., Green, M.R. & Tupler, R. Inappropriate gene activation in FSHD: a repressor complex binds a chromosomal repeat deleted in dystrophic muscle. Cell 110, 339–348 (2002).
El Gazzar, M. et al. Chromatin-specific remodeling by HMGB1 and linker histone H1 silences proinflammatory genes during endotoxin tolerance. Mol. Cell. Biol. 29, 1959–1971 (2009).
Goldfeld, A.E., Birch-Limberger, K., Schooley, R.T. & Walker, B.D. HIV-1 infection does not induce tumor necrosis factor-α or interferon-β gene transcription. J. Acquir. Immune Defic. Syndr. 4, 41–47 (1991).
Haase, A.T. Targeting early infection to prevent HIV-1 mucosal transmission. Nature 464, 217–223 (2010).
Beignon, A.-S. et al. Endocytosis of HIV-1 activates plasmacytoid dendritic cells via Toll-like receptor-viral RNA interactions. J. Clin. Invest. 115, 3265–3275 (2005).
Takaoka, A. et al. DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448, 501–505 (2007).
Siva, A. & Bushman, F. Poly(ADP-ribose) polymerase 1 is not strictly required for infection of murine cells by retroviruses. J. Virol. 76, 11904–11910 (2002).
Pagans, S. et al. SIRT1 regulates HIV transcription via Tat deacetylation. PLoS Biol. 3, e41 (2005).
Fan, Z., Beresford, P., Oh, D., Zhang, D. & Lieberman, J. Tumor suppressor NM23–H1 is a granzyme A-activated DNase during CTL-mediated apoptosis, and the nucleosome assembly protein SET is its inhibitor. Cell 112, 659–672 (2003).
Zandman-Goddard, G. & Shoenfeld, Y. HIV and autoimmunity. Autoimmun. Rev. 1, 329–337 (2002).
Morita, M. et al. Gene-targeted mice lacking the Trex1 (DNase III) 3′→5′ DNA exonuclease develop inflammatory myocarditis. Mol. Cell. Biol. 24, 6719–6727 (2004).
Brass, A.L. et al. Identification of host proteins required for HIV infection through a functional genomic screen. Science 319, 921–926 (2008).
He, J. et al. CCR3 and CCR5 are co-receptors for HIV-1 infection of microglia. Nature 385, 645–649 (1997).
Shun, M.C., Daigle, J.E., Vandegraaff, N. & Engelman, A. Wild-type levels of human immunodeficiency virus type 1 infectivity in the absence of cellular emerin protein. J. Virol. 81, 166–172 (2007).
Hirt, B. Selective extraction of polyoma DNA from infected mouse cell cultures. J. Mol. Biol. 26, 365–369 (1967).
Paz, S. et al. Ubiquitin-regulated recruitment of IκB kinase ɛ to the MAVS interferon signaling adapter. Mol. Cell. Biol. 29, 3401–3412 (2009).
Uzri, D. & Gehrke, L. Nucleotide sequences and modifications that determine RIG-I/RNA binding and signaling activities. J. Virol. 83, 4174–4184 (2009).
Acknowledgements
We thank D. Stetson (University of Washington) for wild-type, Trex1−/− and Trex1−/−Irf3−/− primary MEFs, under agreement with D. Barnes and T. Lindahl (Cancer Research UK); J. Jung (University of Southern California) for RIG-I-deficient (Ddx58−/−) MEFs; S. Harvey and F. Perrino (Wake Forest University) for wild-type and Trex1−/− transformed MEFs; A. Engelman (Dana-Farber Cancer Institute) for the HIV-luciferase plasmid (pNL4-3/Env−); D. Gabuzda (Dana-Farber Cancer Institute) for the HIV-GFP plasmid (pNL4-3/Env−); T. Fujita (Kyoto University) for antiserum to mouse IRF3; S. Nagata (Kyoto University) for the hemagglutinin-tagged MAVS plasmid; K. Fitzgerald (University of Massachusetts) for siRNA and inhibitors of RNA polymerase III; J. Hiscott (McGill University) for IFNB-luciferase plasmid; L. Gehrke (Harvard Medical School) for CMV–renilla luciferase plasmid; and members of the Lieberman lab for discussions. Supported by the US National Institutes of Health (AI45587 to J.L., and T32 HL066987 to N.Y.), the Harvard Center for AIDS Research (N.Y.), Harvard Summer Honors Undergraduate Research Program (A.D.R.-M.) and Deutsche Forschungsgemeinschaft (Le 1074/3-1 to M.A.L.-K.).
Author information
Authors and Affiliations
Contributions
N.Y. conceived of the study, designed and did most experiments and helped write the paper; A.D.R.-M. and B.S. helped do the experiments; M.A.L.-K. provided human cell lines and scientific advice; and J.L. conceived of and supervised the study and helped write the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1–2 (PDF 1971 kb)
Rights and permissions
About this article
Cite this article
Yan, N., Regalado-Magdos, A., Stiggelbout, B. et al. The cytosolic exonuclease TREX1 inhibits the innate immune response to human immunodeficiency virus type 1. Nat Immunol 11, 1005–1013 (2010). https://doi.org/10.1038/ni.1941
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.1941
This article is cited by
-
Genetic determinants of host- and virus-derived insertions for hepatitis E virus replication
Nature Communications (2024)
-
Association of TREX1 polymorphism with disease progression in human immunodeficiency virus type-1 (HIV-1) infected patients
Virus Genes (2023)
-
Evasion of cGAS and TRIM5 defines pandemic HIV
Nature Microbiology (2022)
-
Cytosolic and nuclear recognition of virus and viral evasion
Molecular Biomedicine (2021)
-
Constitutive immune mechanisms: mediators of host defence and immune regulation
Nature Reviews Immunology (2021)