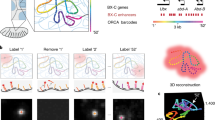
Abstract
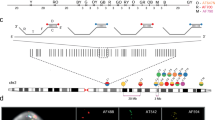
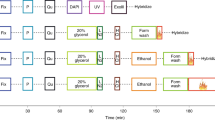
Fluorescent in situ hybridization (FISH) is a powerful, direct and sensitive technique with a wide resolution range that enables the simultaneous study of multiple targets, labelled in different colours. Spreading techniques, denoted here as ‘Fiber-FISH’, increase FISH-resolution to the DMA fiber, using decondensed nuclear DMA as hybridization target1–5. FISH could be a powerful analytical tool for thorough physical examination of yeast artificial chromosomes (YACs) which are often chimaeric or contain internal deletions. However, with one exception restricted to meiotic yeast chromosomes6, FISH has not been used successfully on yeast/YAC DMA. We have developed a fast and simple method that can be applied routinely for compositional and structural analysis of cosmid and YAC DMA in yeast. It enables precise localization and ordering of clones, resolves overlaps and distances and gives a detailed picture of the integrity and colinearity of both probe and target. The combination of high resolution, signal abundance and short yeast cell cycle allows direct visualization of replicating DMA fibers. In a 400 kb region of the human dystrophin gene, we identified two replication origins, demonstrating that human DMA cloned in yeast is capable of initiating its own replication.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Heng, H.H.Q., Squire, J. & Tsui, L.C. High resolution mapping af mammalian genes by in situ hybridization to free chromatin. Proc. natn. Acad. Sci. U.S.A. 89, 9509–9513 (1992).
Wiegant, J. et al. High-resolution in situ hybridization using DNA halo preparations. Hum. molec. Genet. 1, 587–591 (1992).
Parra, I. & Windle, B. High resolution visual mapping of stretched DNA by fluorescent hybridization. Nature Genet. 5, 17–21 (1993).
Haaf, T. & Ward, D.C. High resolution ordering of YAC contigs using extended chromatin and chromosomes. Hum. molec. Genet. 3, 629–633 (1994).
Heiskanen, M., Karhu, R., Hellsten, E., Peltonen, L., Kallioniemi, O.P. & Palotie, A. High resolution mapping using fluorescence in situ hybridization to extended DNA fibers prepared from agarose-embedded cells. Bio Techniques 17, 928–933 (1994).
Scherthan, H., Schweizer, D. & Loidl, J. Delineation of individual chromosomes of Saccharomyces cerevisiae by two-colour in situ hybridization. Trends Genet. 9, 41 (1993).
Florijn, R.J. et al. High-resolution FISH for genomic DNA mapping and colour bar-coding of large genes. Hum. molec. Genet. 4, 831–836 (1995).
Blonden, L.A.J. et al. 242 breakpoints in the 200-kb deletion-prone P20 region of the DMD-gene are widely spread. Genomics 10, 631–639 (1991).
Roth, G.E., Blanton, H.M. & Zakian, V.A. Isolation and characterization of sequences from mouse chromosomal DNA with ARS function in yeast. Molec. cell. Biol. 3, 1898–1908 (1983).
Newlon, C.S. Yeast chromosome replication and segregation. Microbiol. Rev. 52, 568–601 (1988).
Monteil, J.F. et al. Characterization of human chromosomal DNA sequences which replicate autonomously in Saccharomyces cerevisiae. Nucl. Acids Res. 12, 1049–1068 (1984).
Edenberg, H.J. & Huberman, J.A. Eukaryotic DNA replication. Annu. Rev. Genet. 9, 245–284 (1975).
Selig, S., Okumura, K., Ward, D.C. & Cedar, H. Delineation of DNA replication time zones by fluorescence in situ hybridization. EMBO J. 11, 1217–1225 (1992).
Kitsberg, D., Selig, S., Keshet, I. & Cedar, H. Replication structure of the beta-globin gene domain. Nature 366, 588–590 (1993).
Koenig, M. et al. Complete cloning of the Duchenne muscular dystrophy (DMD) cDNA and preliminary genomic organization of the DMD gene in normal and affected individuals. Cell 50, 509–517 (1987).
Den Dunnen, J.T. et al. Reconstruction of the 2.4 Mb human DMD-gene by homologous YAC recombination. Hum. molec. Genet 1, 19–28 (1992).
Kievits, T. et al. Rapid sub-chromosomal localization of cosmids by non-radioactive in situ hybridization. Cytogenet. Cell Genet. 53, 134–136 (1990).
Rights and permissions
About this article
Cite this article
Rosenberg, C., Florijn, R., Van De Rijke, F. et al. High resolution DNA Fiber–fish on yeast artificial chromosomes: direct visualization of DNA replication. Nat Genet 10, 477–479 (1995). https://doi.org/10.1038/ng0895-477
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0895-477
This article is cited by
-
Genomic analysis of a 1 Mb region near the telomere of Hessian fly chromosome X2 and avirulence gene vH13
BMC Genomics (2006)
-
Counting the repetitive kringle-IV repeats in the gene encoding human apolipoprotein(a) by fibre-FISH
Nature Genetics (1999)
-
Replication timing properties, of the humanHPRT locus on active, inactive and reactivated X chromosomes
Somatic Cell and Molecular Genetics (1997)
-
A candidate gene for familial Mediterranean fever
Nature Genetics (1997)