Abstract
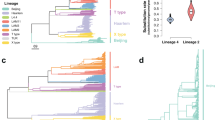
M. tuberculosis is evolving antibiotic resistance, threatening attempts at tuberculosis epidemic control. Mechanisms of resistance, including genetic changes favored by selection in resistant isolates, are incompletely understood. Using 116 newly sequenced and 7 previously sequenced M. tuberculosis whole genomes, we identified genome-wide signatures of positive selection specific to the 47 drug-resistant strains. By searching for convergent evolution—the independent fixation of mutations in the same nucleotide position or gene—we recovered 100% of a set of known resistance markers. We also found evidence of positive selection in an additional 39 genomic regions in resistant isolates. These regions encode components in cell wall biosynthesis, transcriptional regulation and DNA repair pathways. Mutations in these regions could directly confer resistance or compensate for fitness costs associated with resistance. Functional genetic analysis of mutations in one gene, ponA1, demonstrated an in vitro growth advantage in the presence of the drug rifampicin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
NCBI Reference Sequence
Sequence Read Archive
References
World Health Organization. Global Tuberculosis Control 2011 (World Health Organization Press, Geneva, 2011).
Campbell, P.J. et al. Molecular detection of mutations associated with first- and second-line drug resistance compared with conventional drug susceptibility testing of Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 55, 2032–2041 (2011).
Nikaido, H. Prevention of drug access to bacterial targets: permeability barriers and active efflux. Science 264, 382–388 (1994).
Schrag, S.J., Perrot, V. & Levin, B.R. Adaptation to the fitness costs of antibiotic resistance in Escherichia coli. Proc. Biol. Sci. 264, 1287–1291 (1997).
Denamur, E. & Matic, I. Evolution of mutation rates in bacteria. Mol. Microbiol. 60, 820–827 (2006).
Namouchi, A., Didelot, X., Schöck, U., Gicquel, B. & Rocha, E.P.C. After the bottleneck: genome-wide diversification of the Mycobacterium tuberculosis complex by mutation, recombination, and natural selection. Genome Res. 22, 721–734 (2012).
Shapiro, B.J., David, L.A., Friedman, J. & Alm, E.J. Looking for Darwin's footprints in the microbial world. Trends Microbiol. 17, 196–204 (2009).
Kryazhimskiy, S. & Plotkin, J.B. The population genetics of dN/dS. PLoS Genet. 4, e1000304 (2008).
Agashe, D., Martinez-Gomez, N.C., Drummond, D.A. & Marx, C.J. Good codons, bad transcript: large reductions in gene expression and fitness arising from synonymous mutations in a key enzyme. Mol. Biol. Evol. 30, 549–560 (2013).
Rokas, A. & Carroll, S.B. Frequent and widespread parallel evolution of protein sequences. Mol. Biol. Evol. 25, 1943–1953 (2008).
Casali, N. et al. Microevolution of extensively drug-resistant tuberculosis in Russia. Genome Res. 22, 735–745 (2012).
Comas, I. et al. Whole-genome sequencing of rifampicin-resistant Mycobacterium tuberculosis strains identifies compensatory mutations in RNA polymerase genes. Nat. Genet. 44, 106–110 (2012).
Brandis, G., Wrande, M., Liljas, L. & Hughes, D. Fitness-compensatory mutations in rifampicin-resistant RNA polymerase. Mol. Microbiol. 85, 142–151 (2012).
de Vos, M. et al. Putative compensatory mutations in the rpoC gene of rifampin-resistant Mycobacterium tuberculosis are associated with ongoing transmission. Antimicrob. Agents Chemother. 57, 827–832 (2013).
Tanabe, K., Kondo, T., Onodera, Y. & Furusawa, M. A conspicuous adaptability to antibiotics in the Escherichia coli mutator strain, dnaQ49. FEMS Microbiol. Lett. 176, 191–196 (1999).
Dos Vultos, T., Mestre, O., Tonjum, T. & Gicquel, B. DNA repair in Mycobacterium tuberculosis revisited. FEMS Microbiol. Rev. 33, 471–487 (2009).
Soldini, S. et al. PPE_MPTR genes are differentially expressed by Mycobacterium tuberculosis in vivo. Tuberculosis (Edinb.) 91, 563–568 (2011).
Kaur, D., Guerin, M.E., Skovierová, H., Brennan, P.J. & Jackson, M. Chapter 2: biogenesis of the cell wall and other glycoconjugates of Mycobacterium tuberculosis. Adv. Appl. Microbiol. 69, 23–78 (2009).
Yu, J. et al. Both phthiocerol dimycocerosates and phenolic glycolipids are required for virulence of Mycobacterium marinum. Infect. Immun. 80, 1381–1389 (2012).
Matsunaga, I. et al. Mycobacterium tuberculosis pks12 produces a novel polyketide presented by CD1c to T cells. J. Exp. Med. 200, 1559–1569 (2004).
Dubey, V.S., Sirakova, T.D. & Kolattukudy, P.E. Disruption of msl3 abolishes the synthesis of mycolipanoic and mycolipenic acids required for polyacyltrehalose synthesis in Mycobacterium tuberculosis H37Rv and causes cell aggregation. Mol. Microbiol. 45, 1451–1459 (2002).
Hett, E.C., Chao, M.C. & Rubin, E.J. Interaction and modulation of two antagonistic cell wall enzymes of mycobacteria. PLoS Pathog. 6, e1001020 (2010).
Billman-Jacobe, H., Haites, R.E. & Coppel, R.L. Characterization of a Mycobacterium smegmatis mutant lacking penicillin binding protein 1. Antimicrob. Agents Chemother. 43, 3011–3013 (1999).
Philalay, J.S., Palermo, C.O., Hauge, K.A., Rustad, T.R. & Cangelosi, G.A. Genes required for intrinsic multidrug resistance in Mycobacterium avium. Antimicrob. Agents Chemother. 48, 3412–3418 (2004).
Chavadi, S.S. et al. Inactivation of tesA reduces cell wall lipid production and increases drug susceptibility in mycobacteria. J. Biol. Chem. 286, 24616–24625 (2011).
Matsunaga, I., Meda, S., Nakata, N. & Fujiwara, N. The polyketide synthase–associated multidrug tolerance in Mycobacterium intracellulare clinical isolates. Chemotherapy 58, 341–348 (2012).
Bisson, G.P. et al. Upregulation of the phthiocerol dimycocerosate biosynthetic pathway by rifampin-resistant, rpoB mutant Mycobacterium tuberculosis. J. Bacteriol. 194, 6441–6452 (2012).
Sun, G. et al. Dynamic population changes in Mycobacterium tuberculosis during acquisition and fixation of drug resistance in patients. J. Infect. Dis. 206, 1724–1733 (2012).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Shigemura, K. et al. Presence of a mutation in ponA1 of Neisseria gonorrhoeae in numerous clinical samples resistant to various β-lactams and other, structurally unrelated, antimicrobials. J. Infect. Chemother. 11, 226–230 (2005).
Zahrt, T.C. & Deretic, V. An essential two-component signal transduction system in Mycobacterium tuberculosis. J. Bacteriol. 182, 3832–3838 (2000).
Nguyen, H.T., Wolff, K.A., Cartabuke, R.H., Ogwang, S. & Nguyen, L. A lipoprotein modulates activity of the MtrAB two-component system to provide intrinsic multidrug resistance, cytokinetic control and cell wall homeostasis in Mycobacterium. Mol. Microbiol. 76, 348–364 (2010).
Cangelosi, G.A. et al. The two-component regulatory system mtrAB is required for morphotypic multidrug resistance in Mycobacterium avium. Antimicrob. Agents Chemother. 50, 461–468 (2006).
Möker, N. et al. Deletion of the genes encoding the MtrA-MtrB two-component system of Corynebacterium glutamicum has a strong influence on cell morphology, antibiotics susceptibility and expression of genes involved in osmoprotection. Mol. Microbiol. 54, 420–438 (2004).
Lew, J.M., Kapopoulou, A., Jones, L.M. & Cole, S.T. TubercuList—10 years after. Tuberculosis (Edinb.) 91, 1–7 (2011).
Jiang, X. et al. Comparison of the proteome of isoniazid-resistant and -susceptible strains of Mycobacterium tuberculosis. Microb. Drug Resist. 12, 231–238 (2006).
Yang, Q., Liu, Y., Huang, F. & He, Z.-G. Physical and functional interaction between D-ribokinase and topoisomerase I has opposite effects on their respective activity in Mycobacterium smegmatis and Mycobacterium tuberculosis. Arch. Biochem. Biophys. 512, 135–142 (2011).
Sandgren, A. et al. Tuberculosis drug resistance mutation database. PLoS Med. 6, e2 (2009).
Nessar, R., Reyrat, J.M., Murray, A. & Gicquel, B. Genetic analysis of new 16S rRNA mutations conferring aminoglycoside resistance in Mycobacterium abscessus. J. Antimicrob. Chemother. 66, 1719–1724 (2011).
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008).
Comas, I. et al. Human T cell epitopes of Mycobacterium tuberculosis are evolutionarily hyperconserved. Nat. Genet. 42, 498–503 (2010).
Felsenstein, J. PHYLIP—Phylogeny Inference Package (Version 3.2). Cladistics 5, 164–166 (1989).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012).
Guindon, S. et al. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst. Biol. 59, 307–321 (2010).
Popescu, A.-A., Huber, K.T. & Paradis, E. ape 3.0: new tools for distance based phylogenetics and evolutionary analysis in R. Bioinformatics 28, 1536–1537 (2012).
Acknowledgements
We thank the technical staff of the British Columbia Centre for Disease Control Public Health Microbiology and Reference Mycobacteriology Laboratory in Vancouver, M. Bosman from the National Health Laboratory Service in Cape Town and L. Fattorini from the Istituto Superiore di Sanita in Rome. This work was funded by a Senior Ellison Foundation Award (M.M.) and in part by a contact from the National Institute of Allergy and Infectious Diseases (HHSN266200400001C to B.B.), the Department of Pulmonary and Critical Care at Massachusetts General Hospital (M.R.F.), a postdoctoral fellowship from the Harvard MIDAS Center for Communicable Disease Dynamics (B.J.S.) and a Packard Foundation Fellowship (P.C.S.). S.G. was supported by the Swiss National Science Foundation (PP0033_119205).
Author information
Authors and Affiliations
Contributions
This study was designed and conducted by M.R.F. and M.M. M.R.F. wrote the first drafts of the manuscript. B.J.S., P.C.S. and E.S.L. provided conceptual input on the evolutionary testing, analysis support and key manuscript edits. K.J.K. and E.J.R. constructed the ponA1 mutants and measured their MICs. R.S. provided bioinformatics support and K.R.J. helped with the curation of the isolate phenotypes. R.M.W., E.M.S., T.C.V. and A.C. conducted molecular epidemiological studies and performed molecular characterization, drug susceptibility testing and selection of isolates from South Africa. A.S. and D.K. performed molecular characterization and drug sensitivity testing and selected isolates from Peru and Russia. B.P. and J.E.P. performed molecular characterization, drug sensitivity testing and selection of isolates from the Centers for Disease Control and Prevention. M.R.O. identified the individual with progressively resistant tuberculosis and performed molecular characterization and selection of serial isolates from this individual in Italy. J.L.G., J.C.J., M.R. and P.K.C.T. conducted the tuberculosis outbreak investigation in British Columbia and performed molecular characterization, drug susceptibility testing and sequencing of these isolates. M.K.-M. conducted the epidemiological study of tuberculosis transmission in San Francisco, and M.L.B. and B.M. performed molecular characterization and sequencing of these isolates. B.N.K. and N.K. characterized the W-148, Haarlem and C isolates. S.G. collected the 24 drug-sensitive M. tuberculosis diversity strain set. J.G. and B.B. provided oversight for sequencing and bioinformatics support.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–10 and 12–20, and Supplementary Note (PDF 7052 kb)
Supplementary Table 11
Description of the M. tuberculosis isolates studied (XLSX 47 kb)
Supplementary Table 21
Multiple alignment of nucleotide sequence at all variable sites within the targets of independent mutation (XLSX 243 kb)
Source data
Rights and permissions
About this article
Cite this article
Farhat, M., Shapiro, B., Kieser, K. et al. Genomic analysis identifies targets of convergent positive selection in drug-resistant Mycobacterium tuberculosis. Nat Genet 45, 1183–1189 (2013). https://doi.org/10.1038/ng.2747
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2747
This article is cited by
-
Detecting patterns of accessory genome coevolution in Staphylococcus aureus using data from thousands of genomes
BMC Bioinformatics (2023)
-
Statistical analysis of 914 Mycobacterium tuberculosis genomes reveals single nucleotide polymorphisms in the ponA1 gene associated with rifampicin resistance
The Nucleus (2023)
-
Population genomics of Group B Streptococcus reveals the genetics of neonatal disease onset and meningeal invasion
Nature Communications (2022)
-
PowerBacGWAS: a computational pipeline to perform power calculations for bacterial genome-wide association studies
Communications Biology (2022)
-
Reversible gene silencing through frameshift indels and frameshift scars provide adaptive plasticity for Mycobacterium tuberculosis
Nature Communications (2021)