Abstract
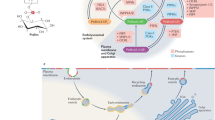
Phosphoinositide (PI) lipids are essential components of eukaryotic cell membranes. They are produced by mono-, bis- and trisphosphorylation of the inositol headgroup of phosphatidylinositol (PtdIns) and are concentrated in separate pools of cytosolic membranes. PIs serve as markers of the cell compartments and form unique docking sites for protein effectors. Collectively, seven known PIs, the protein effectors that bind them and enzymes that generate or modify PIs compose a remarkably complex protein-lipid signaling network. A number of cytosolic proteins contain one or several effector modules capable of recognizing individual PIs and recruiting the host proteins to distinct intracellular compartment. The recently determined atomic-resolution structures and membrane-targeting mechanisms of a dozen PI effectors have provided insights into the molecular basis for regulation of endocytic membrane trafficking and signaling. In this review, I highlight the structural aspects of the deciphering of the 'PI code' by the most common PI-recognizing effectors and discuss the mechanistic details of their membrane anchoring.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lemmon, M.A. Membrane recognition by phospholipid-binding domains. Nat. Rev. Mol. Cell Biol. 9, 99–111 (2008).
Hurley, J.H. Membrane binding domains. Biochim Biophys Acta 1761, 805–811 (2006).
Di Paolo, G. & De Camilli, P. Phosphoinositides in cell regulation and membrane dynamics. Nature 443, 651–657 (2006).
Roth, M.G. Phosphoinositides in constitutive membrane traffic. Physiol. Rev. 84, 699–730 (2004).
Vanhaesebroeck, B. et al. Synthesis and function of 3-phosphorylated inositol lipids. Annu. Rev. Biochem. 70, 535–602 (2001).
Harlan, J.E., Hajduk, P.J., Yoon, H.S. & Fesik, S.W. Pleckstrin homology domains bind to phosphatidylinositol-4,5-bisphosphate. Nature 371, 168–170 (1994).
Kutateladze, T.G. Phosphatidylinositol 3-phosphate recognition and membrane docking by the FYVE domain. Biochim. Biophys. Acta 1761, 868–877 (2006).
Burd, C.G. & Emr, S.D. Phosphatidylinositol(3)-phosphate signaling mediated by specific binding to RING FYVE domains. Mol. Cell 2, 157–162 (1998).
Gaullier, J.M. et al. FYVE fingers bind PtdIns(3)P. Nature 394, 432–433 (1998).
Patki, V., Lawe, D.C., Corvera, S., Virbasius, J.V. & Chawla, A. A functional PtdIns(3)P-binding motif. Nature 394, 433–434 (1998).
Ridley, S.H. et al. FENS-1 and DFCP1 are FYVE domain–containing proteins with distinct functions in the endosomal and Golgi compartments. J. Cell Sci. 114, 3991–4000 (2001).
Simonsen, A. et al. Alfy, a novel FYVE-domain-containing protein associated with protein granules and autophagic membranes. J. Cell Sci. 117, 4239–4251 (2004).
Gillooly, D.J., Simonsen, A. & Stenmark, H. Cellular functions of phosphatidylinositol 3-phosphate and FYVE domain proteins. Biochem. J. 355, 249–258 (2001).
Stahelin, R.V. et al. Phosphatidylinositol 3-phosphate induces the membrane penetration of the FYVE domains of Vps27p and Hrs. J. Biol. Chem. 277, 26379–26388 (2002).
Diraviyam, K., Stahelin, R.V., Cho, W. & Murray, D. Computer modeling of the membrane interaction of FYVE domains. J. Mol. Biol. 328, 721–736 (2003).
Kutateladze, T.G. et al. Multivalent mechanism of membrane insertion by the FYVE domain. J. Biol. Chem. 279, 3050–3057 (2004).
Lee, S.A. et al. Targeting of the FYVE domain to endosomal membranes is regulated by a histidine switch. Proc. Natl. Acad. Sci. USA 102, 13052–13057 (2005).
He, J. et al. Membrane insertion of the FYVE domain is modulated by pH. Proteins 76, 852–860 (2009).
Mertens, H.D., Callaghan, J.M., Swarbrick, J.D., McConville, M.J. & Gooley, P.R. A high-resolution solution structure of a trypanosomatid FYVE domain. Protein Sci. 16, 2552–2559 (2007).
Misra, S. & Hurley, J.H. Crystal structure of a phosphatidylinositol 3-phosphate-specific membrane-targeting motif, the FYVE domain of Vps27p. Cell 97, 657–666 (1999).
Kutateladze, T. & Overduin, M. Structural mechanism of endosome docking by the FYVE domain. Science 291, 1793–1796 (2001).
Blatner, N.R. et al. The molecular basis of the differential subcellular localization of FYVE domains. J. Biol. Chem. 279, 53818–53827 (2004).
Sankaran, V.G., Klein, D.E., Sachdeva, M.M. & Lemmon, M.A. High affinity binding of a FYVE domain to phosphatidylinositol 3-phosphate requires intact phospholipid but not FYVE domain oligomerization. Biochemistry 40, 8581–8587 (2001).
Brunecky, R. et al. Investigation of the binding geometry of a peripheral membrane protein. Biochemistry 44, 16064–16071 (2005).
Callaghan, J., Simonsen, A., Gaullier, J.M., Toh, B.H. & Stenmark, H. The endosome fusion regulator early-endosomal autoantigen 1 (EEA1) is a dimer. Biochem. J. 338, 539–543 (1999).
Dumas, J.J. et al. Multivalent endosome targeting by homodimeric EEA1. Mol. Cell 8, 947–958 (2001).
Lawe, D.C., Patki, V., Heller-Harrison, R., Lambright, D. & Corvera, S. The FYVE domain of early endosome antigen 1 is required for both phosphatidylinositol 3-phosphate and Rab5 binding. Critical role of this dual interaction for endosomal localization. J. Biol. Chem. 275, 3699–3705 (2000).
Hayakawa, A. et al. Structural basis for endosomal targeting by FYVE domains. J. Biol. Chem. 279, 5958–5966 (2004).
Mao, Y. et al. Crystal structure of the VHS and FYVE tandem domains of Hrs, a protein involved in membrane trafficking and signal transduction. Cell 100, 447–456 (2000).
Lemmon, M.A. & Ferguson, K.M. Signal-dependent membrane targeting by pleckstrin homology (PH) domains. Biochem. J. 350, 1–18 (2000).
DiNitto, J.P. & Lambright, D.G. Membrane and juxtamembrane targeting by PH and PTB domains. Biochim. Biophys. Acta 1761, 850–867 (2006).
Haslam, R.J., Koide, H.B. & Hemmings, B.A. Pleckstrin domain homology. Nature 363, 309–310 (1993).
Mayer, B.J., Ren, R., Clark, K.L. & Baltimore, D. A putative modular domain present in diverse signaling proteins. Cell 73, 629–630 (1993).
Fukuda, M., Kojima, T., Kabayama, H. & Mikoshiba, K. Mutation of the pleckstrin homology domain of Bruton's tyrosine kinase in immunodeficiency impaired inositol 1,3,4,5-tetrakisphosphate binding capacity. J. Biol. Chem. 271, 30303–30306 (1996).
Klarlund, J.K. et al. Signaling by phosphoinositide-3,4,5-trisphosphate through proteins containing pleckstrin and Sec7 homology domains. Science 275, 1927–1930 (1997).
Cronin, T.C., DiNitto, J.P., Czech, M.P. & Lambright, D.G. Structural determinants of phosphoinositide selectivity in splice variants of Grp1 family PH domains. EMBO J. 23, 3711–3720 (2004).
Lemmon, M.A. Pleckstrin homology domains: not just for phosphoinositides. Biochem. Soc. Trans. 32, 707–711 (2004).
Singh, S.M. & Murray, D. Molecular modeling of the membrane targeting of phospholipase C pleckstrin homology domains. Protein Sci. 12, 1934–1953 (2003).
Manna, D., Albanese, A., Park, W.S. & Cho, W. Mechanistic basis of differential cellular responses of phosphatidylinositol 3,4-bisphosphate- and phosphatidylinositol 3,4,5-trisphosphate-binding pleckstrin homology domains. J. Biol. Chem. 282, 32093–32105 (2007).
He, J. et al. Molecular mechanism of membrane targeting by the GRP1 PH domain. J. Lipid Res. 49, 1807–1815 (2008).
Knight, J.D. & Falke, J.J. Single-molecule fluorescence studies of a PH domain: new insights into the membrane docking reaction. Biophys. J. 96, 566–582 (2009).
Ferguson, K.M. et al. Structural basis for discrimination of 3-phosphoinositides by pleckstrin homology domains. Mol. Cell 6, 373–384 (2000).
Lietzke, S.E. et al. Structural basis of 3-phosphoinositide recognition by pleckstrin homology domains. Mol. Cell 6, 385–394 (2000).
Flesch, F.M., Yu, J.W., Lemmon, M.A. & Burger, K.N. Membrane activity of the phospholipase C-delta1 pleckstrin homology (PH) domain. Biochem. J. 389, 435–441 (2005).
Stahelin, R.V., Karathanassis, D., Murray, D., Williams, R.L. & Cho, W. Structural and membrane binding analysis of the Phox homology domain of Bem1p: basis of phosphatidylinositol 4-phosphate specificity. J. Biol. Chem. 282, 25737–25747 (2007).
Komander, D. et al. Structural insights into the regulation of PDK1 by phosphoinositides and inositol phosphates. EMBO J. 23, 3918–3928 (2004).
Baraldi, E. et al. Structure of the PH domain from Bruton′s tyrosine kinase in complex with inositol 1,3,4,5-tetrakisphosphate. Structure 7, 449–460 (1999).
DiNitto, J.P. et al. Structural basis and mechanism of autoregulation in 3-phosphoinositide-dependent Grp1 family Arf GTPase exchange factors. Mol. Cell 28, 569–583 (2007).
Thomas, C.C., Deak, M., Alessi, D.R. & van Aalten, D.M. High-resolution structure of the pleckstrin homology domain of protein kinase b/akt bound to phosphatidylinositol (3,4,5)-trisphosphate. Curr. Biol. 12, 1256–1262 (2002).
Milburn, C.C. et al. Binding of phosphatidylinositol 3,4,5-trisphosphate to the pleckstrin homology domain of protein kinase B induces a conformational change. Biochem. J. 375, 531–538 (2003).
Carpten, J.D. et al. A transforming mutation in the pleckstrin homology domain of AKT1 in cancer. Nature 448, 439–444 (2007).
Ferguson, K.M., Lemmon, M.A., Schlessinger, J. & Sigler, P.B. Structure of the high affinity complex of inositol trisphosphate with a phospholipase C pleckstrin homology domain. Cell 83, 1037–1046 (1995).
Jackson, S.G., Zhang, Y., Haslam, R.J. & Junop, M.S. Structural analysis of the carboxy terminal PH domain of pleckstrin bound to D-myo-inositol 1,2,3,5,6-pentakisphosphate. BMC Struct. Biol. 7, 80 (2007).
Ceccarelli, D.F. et al. Non-canonical interaction of phosphoinositides with pleckstrin homology domains of Tiam1 and ArhGAP9. J. Biol. Chem. 282, 13864–13874 (2007).
Hyvönen, M. et al. Structure of the binding site for inositol phosphates in a PH domain. EMBO J. 14, 4676–4685 (1995).
Seet, L.F. & Hong, W. The Phox (PX) domain proteins and membrane traffic. Biochim. Biophys. Acta 1761, 878–896 (2006).
Ponting, C.P. et al. Novel domains in NADPH oxidase subunits, sorting nexins, and PtdIns 3-kinases: binding partners of SH3 domains? Protein Sci. 5, 2353–2357 (1996).
Yu, J.W. & Lemmon, M.A. All phox homology (PX) domains from Saccharomyces cerevisiae specifically recognize phosphatidylinositol-3-phosphate. J. Biol. Chem. 276, 44179–44184 (2001).
Kanai, F. et al. The PX domains of p47phox and p40phox bind to lipid products of PI(3)K. Nat. Cell Biol. 3, 675–678 (2001).
Xu, Y., Hortsman, H., Seet, L., Wong, S.H. & Hong, W. SNX3 regulates endosomal function through its PX-domain-mediated interaction with PtdIns(3)P. Nat. Cell Biol. 3, 658–666 (2001).
Cheever, M.L. et al. Phox domain interaction with PtdIns(3)P targets the Vam7 t-SNARE to vacuole membranes. Nat. Cell Biol. 3, 613–618 (2001).
Ellson, C.D. et al. PtdIns(3)P regulates the neutrophil oxidase complex by binding to the PX domain of p40(phox). Nat. Cell Biol. 3, 679–682 (2001).
Song, X. et al. Phox homology domains specifically bind phosphatidylinositol phosphates. Biochemistry 40, 8940–8944 (2001).
Málková, S., Stahelin, R.V., Pingali, S.V., Cho, W. & Schlossman, M.L. Orientation and penetration depth of monolayer-bound p40phox-PX. Biochemistry 45, 13566–13575 (2006).
Stahelin, R.V., Burian, A., Bruzik, K.S., Murray, D. & Cho, W. Membrane binding mechanisms of the PX domains of NADPH oxidase p40phox and p47phox. J. Biol. Chem. 278, 14469–14479 (2003).
Lee, S.A. et al. Molecular mechanism of membrane docking by the Vam7p PX domain. J. Biol. Chem. 281, 37091–37101 (2006).
Bravo, J. et al. The crystal structure of the PX domain from p40(phox) bound to phosphatidylinositol 3-phosphate. Mol. Cell 8, 829–839 (2001).
Karathanassis, D. et al. Binding of the PX domain of p47(phox) to phosphatidylinositol 3,4-bisphosphate and phosphatidic acid is masked by an intramolecular interaction. EMBO J. 21, 5057–5068 (2002).
Zhou, C.Z. et al. Crystal structure of the yeast Phox homology (PX) domain protein Grd19p complexed to phosphatidylinositol-3-phosphate. J. Biol. Chem. 278, 50371–50376 (2003).
Lu, J., Garcia, J., Dulubova, I., Sudhof, T.C. & Rizo, J. Solution structure of the Vam7p PX domain. Biochemistry 41, 5956–5962 (2002).
Hiroaki, H., Ago, T., Ito, T., Sumimoto, H. & Kohda, D. Solution structure of the PX domain, a target of the SH3 domain. Nat. Struct. Biol. 8, 526–530 (2001).
Xing, Y. et al. Structural basis of membrane targeting by the Phox homology domain of cytokine-independent survival kinase (CISK-PX). J. Biol. Chem. 279, 30662–30669 (2004).
Kutateladze, T.G. Mechanistic similarities in docking of the FYVE and PX domains to phosphatidylinositol 3-phosphate containing membranes. Prog. Lipid Res. 46, 315–327 (2007).
Stahelin, R.V. et al. Mechanism of membrane binding of the phospholipase D1 PX domain. J. Biol. Chem. 279, 54918–54926 (2004).
Stahelin, R.V. et al. Structural and membrane binding analysis of the Phox homology domain of phosphoinositide 3-kinase-C2alpha. J. Biol. Chem. 281, 39396–39406 (2006).
Pylypenko, O., Lundmark, R., Rasmuson, E., Carlsson, S.R. & Rak, A. The PX-BAR membrane-remodeling unit of sorting nexin 9. EMBO J. 26, 4788–4800 (2007).
Blatner, N.R. et al. The structural basis of novel endosome anchoring activity of KIF16B kinesin. EMBO J. 26, 3709–3719 (2007).
Honbou, K. et al. Full-length p40phox structure suggests a basis for regulation mechanism of its membrane binding. EMBO J. 26, 1176–1186 (2007).
Parkinson, G.N., Vines, D., Driscoll, P.C. & Djordjevic, S. Crystal structures of PI3K–C2alpha PX domain indicate conformational change associated with ligand binding. BMC Struct. Biol. 8, 13 (2008).
Zhong, Q. et al. Determinants of the endosomal localization of sorting nexin 1. Mol. Biol. Cell 16, 2049–2057 (2005).
Song, J., Zhao, K.Q., Newman, C.L., Vinarov, D.A. & Markley, J.L. Solution structure of human sorting nexin 22. Protein Sci. 16, 807–814 (2007).
Itoh, T. & De Camilli, P. BAR, F-BAR (EFC) and ENTH/ANTH domains in the regulation of membrane-cytosol interfaces and membrane curvature. Biochim. Biophys. Acta 1761, 897–912 (2006).
Cho, W. & Stahelin, R.V. Membrane binding and subcellular targeting of C2 domains. Biochim. Biophys. Acta 1761, 838–849 (2006).
Hamada, K., Shimizu, T., Matsui, T., Tsukita, S. & Hakoshima, T. Structural basis of the membrane-targeting and unmasking mechanisms of the radixin FERM domain. EMBO J. 19, 4449–4462 (2000).
Dippold, H.C. et al. GOLPH3 bridges phosphatidylinositol-4- phosphate and actomyosin to stretch and shape the Golgi to promote budding. Cell 139, 337–351 (2009).
Wood, C.S. et al. PtdIns4P recognition by Vps74/GOLPH3 links PtdIns 4-kinase signaling to retrograde Golgi trafficking. J. Cell Biol. 187, 967–975 (2009).
Zimmermann, P. The prevalence and significance of PDZ domain-phosphoinositide interactions. Biochim. Biophys. Acta 1761, 947–956 (2006).
Dove, S.K., Dong, K., Kobayashi, T., Williams, F.K. & Michell, R.H. Phosphatidylinositol 3,5-bisphosphate and Fab1p/PIKfyve underPPIn endo-lysosome function. Biochem. J. 419, 1–13 (2009).
Santagata, S. et al. G-protein signaling through tubby proteins. Science 292, 2041–2050 (2001).
Szentpetery, Z., Balla, A., Kim, Y.J., Lemmon, M.A. & Balla, T. Live cell imaging with protein domains capable of recognizing phosphatidylinositol 4,5-bisphosphate; a comparative study. BMC Cell Biol. 10, 67 (2009).
Wang, J., Arbuzova, A., Hangyas-Mihalyne, G. & McLaughlin, S. The effector domain of myristoylated alanine-rich C kinase substrate binds strongly to phosphatidylinositol 4,5-bisphosphate. J. Biol. Chem. 276, 5012–5019 (2001).
Caroni, P. New EMBO members' review: actin cytoskeleton regulation through modulation of PI(4,5)P(2) rafts. EMBO J. 20, 4332–4336 (2001).
Dell'Acqua, M.L., Faux, M.C., Thorburn, J., Thorburn, A. & Scott, J.D. Membrane-targeting sequences on AKAP79 bind phosphatidylinositol-4, 5-bisphosphate. EMBO J. 17, 2246–2260 (1998).
Goldschmidt-Clermont, P.J., Machesky, L.M., Baldassare, J.J. & Pollard, T.D. The actin-binding protein profilin binds to PIP2 and inhibits its hydrolysis by phospholipase C. Science 247, 1575–1578 (1990).
Rohatgi, R., Ho, H.Y. & Kirschner, M.W. Mechanism of N-WASP activation by CDC42 and phosphatidylinositol 4, 5-bisphosphate. J. Cell Biol. 150, 1299–1310 (2000).
Ford, M.G. et al. Simultaneous binding of PtdIns(4,5)P2 and clathrin by AP180 in the nucleation of clathrin lattices on membranes. Science 291, 1051–1055 (2001).
Ford, M.G. et al. Curvature of clathrin-coated pits driven by epsin. Nature 419, 361–366 (2002).
McCrea, H.J. & De Camilli, P. Mutations in phosphoinositide metabolizing enzymes and human disease. Physiology (Bethesda) 24, 8–16 (2009).
Prestwich, G.D. Phosphoinositide signaling; from affinity probes to pharmaceutical targets. Chem. Biol. 11, 619–637 (2004).
Engelman, J.A., Luo, J. & Cantley, L.C. The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat. Rev. Genet. 7, 606–619 (2006).
Acknowledgements
The research in T.G.K.'s laboratory is supported by the US National Institutes of Health grants GM 071424 and CA 113472.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Rights and permissions
About this article
Cite this article
Kutateladze, T. Translation of the phosphoinositide code by PI effectors. Nat Chem Biol 6, 507–513 (2010). https://doi.org/10.1038/nchembio.390
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.390