Abstract
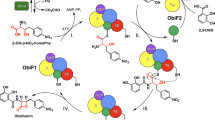
Biosynthetic modification of nonribosomal peptide backbones represents a potentially powerful strategy to modulate the structure and properties of an important class of therapeutics. Using a high-throughput assay for catalytic activity, we show here that an L-Phe-specific module of an archetypal nonribosomal peptide synthetase can be reprogrammed to accept and process the backbone-modified amino acid (S)-β-Phe with near-native specificity and efficiency. A co-crystal structure with a non-hydrolysable aminoacyl-AMP analogue reveals the origins of the 40,000-fold α/β-specificity switch, illuminating subtle but precise remodelling of the active site. When the engineered catalyst was paired with downstream module(s), (S)-β-Phe-containing peptides were produced at preparative scale in vitro (~1 mmol) and high titres in vivo (~100 mg l–1), highlighting the potential of biosynthetic pathway engineering for the construction of novel nonribosomal β-frameworks.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Newman, D. J. & Cragg, G. M. Natural products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 79, 629–661 (2016).
Cane, D. E., Walsh, C. T. & Khosla, C. Harnessing the biosynthetic code: combinations, permutations, and mutations. Science 282, 63–68 (1998).
Kim, E., Moore, B. S. & Yoon, Y. J. Reinvigorating natural product combinatorial biosynthesis with synthetic biology. Nat. Chem. Biol. 11, 649–659 (2015).
Schwarzer, D., Finking, R. & Marahiel, M. A. Nonribosomal peptides: from genes to products. Nat. Prod. Rep. 20, 275–287 (2003).
Felnagle, E. A. et al. Nonribosomal peptide synthetases involved in the production of medically relevant natural products. Mol. Pharm. 5, 191–211 (2008).
Süssmuth, R. D. & Mainz, A. Nonribosomal peptide synthesis—principles and prospects. Angew. Chem. Int. Ed. 56, 3770–3821 (2017).
Sieber, S. A. & Marahiel, M. A. Molecular mechanisms underlying nonribosomal peptide synthesis: approaches to new antibiotics. Chem. Rev. 105, 715–738 (2005).
Fischbach, M. A. & Walsh, C. T. Assembly-line enzymology for polyketide and nonribosomal peptide antibiotics: logic, machinery, and mechanisms. Chem. Rev. 106, 3468–3496 (2006).
Hur, G. H., Vickery, C. R. & Burkart, M. D. Explorations of catalytic domains in non-ribosomal peptide synthetase enzymology. Nat. Prod. Rep. 29, 1074–1098 (2012).
Winn, M., Fyans, J. K., Zhuo, Y. & Micklefield, J. Recent advances in engineering nonribosomal peptide assembly lines. Nat. Prod. Rep. 33, 317–347 (2016).
Eppelmann, K., Stachelhaus, T. & Marahiel, M. A. Exploitation of the selectivity-conferring code of nonribosomal peptide synthetases for the rational design of novel peptide antibiotics. Biochemistry 41, 9718–9726 (2002).
Chen, C.-Y., Georgiev, I., Anderson, A. C. & Donald, B. R. Computational structure-based redesign of enzyme activity. Proc. Natl Acad. Sci. USA 106, 3764–3769 (2009).
Evans, B. S., Chen, Y., Metcalf, W. W., Zhao, H. & Kelleher, N. L. Directed evolution of the nonribosomal peptide synthetase AdmK generates new andrimid derivatives in vivo. Chem. Biol. 18, 601–607 (2011).
Villiers, B. & Hollfelder, F. Directed evolution of a gatekeeper domain in nonribosomal peptide synthesis. Chem. Biol. 18, 1290–1299 (2011).
Thirlway, J. et al. Introduction of a non-natural amino acid into a nonribosomal peptide antibiotic by modification of adenylation domain specificity. Angew. Chem. Int. Ed. 51, 7181–7184 (2012).
Zhang, K. et al. Engineering the substrate specificity of the DhbE adenylation domain by yeast cell surface display. Chem. Biol. 20, 92–101 (2013).
Kries, H. et al. Reprogramming nonribosomal peptide synthetases for ‘clickable’ amino acids. Angew. Chem. Int. Ed. 53, 10105–10108 (2014).
Kaljunen, H. et al. Structural elucidation of the bispecificity of A domains as a basis for activating non-natural amino acids. Angew. Chem. Int. Ed. 54, 8833–8836 (2015).
Shrestha, S. K. & Garneau-Tsodikova, S. Expanding substrate promiscuity by engineering a novel adenylating-methylating NRPS bifunctional enzyme. ChemBioChem 17, 1328–1332 (2016).
Kudo, F., Miyanaga, A. & Eguchi, T. Biosynthesis of natural products containing β-amino acids. Nat. Prod. Rep. 31, 1056–1073 (2014).
Seebach, D. et al. New open-chain and cyclic tetrapeptides, consisting of α-, β2-, and β3-amino-acid residues, as somatostatin mimics—a survey. Helv. Chim. Acta 91, 1736–1786 (2008).
Cheloha, R. W., Maeda, A., Dean, T., Gardella, T. J. & Gellman, S. H. Backbone modification of a polypeptide drug alters duration of action in vivo. Nat. Biotechnol. 32, 653–655 (2014).
Werner, H. M. & Horne, W. S. Folding and function in α/β-peptides: targets and therapeutic applications. Curr. Opin. Chem. Biol. 28, 75–82 (2015).
Mootz, H. D. & Marahiel, M. A. The tyrocidine biosynthesis operon of Bacillus brevis: complete nucleotide sequence and biochemical characterization of functional internal adenylation domains. J. Bacteriol. 179, 6843–6850 (1997).
Boder, E. T. & Wittrup, K. D. Yeast surface display for screening combinatorial polypeptide libraries. Nat. Biotechnol. 15, 553–557 (1997).
Tornøe, C. W., Christensen, C. & Meldal, M. Peptidotriazoles on solid phase: [1,2,3]-triazoles by regiospecific copper(I)-catalysed 1,3-dipolar cycloadditions of terminal alkynes to azides. J. Org. Chem. 67, 3057–3064 (2002).
Rostovtsev, V. V., Green, L. G., Fokin, V. V. & Sharpless, K. B. A stepwise Huisgen cycloaddition process: copper(I)-catalysed regioselective ‘ligation’ of azides and terminal alkynes. Angew. Chem. Int. Ed. 41, 2596–2599 (2002).
Conti, E., Stachelhaus, T., Marahiel, M. A. & Brick, P. Structural basis for the activation of phenylalanine in the non-ribosomal biosynthesis of gramicidin S. EMBO J. 16, 4174–4183 (1997).
Miyanaga, A., Cieślak, J., Shinohara, Y., Kudo, F. & Eguchi, T. The crystal structure of the adenylation enzyme VinN reveals a unique β-amino acid recognition mechanism. J. Biol. Chem. 289, 31448–31457 (2014).
Miyanaga, A., Kudo, F. & Eguchi, T. Mechanisms of β-amino acid incorporation in polyketide macrolactam biosynthesis. Curr. Opin. Chem. Biol. 35, 58–64 (2016).
Otten, L. G., Schaffer, M. L., Villiers, B. R. M., Stachelhaus, T. & Hollfelder, F. An optimized ATP/PPi-exchange assay in 96-well format for screening of adenylation domains for applications in combinatorial biosynthesis. Biotechnol. J. 2, 232–240 (2007).
Stachelhaus, T., Mootz, H. D., Bergendahl, V. & Marahiel, M. A. Peptide bond formation in nonribosomal peptide biosynthesis. Catalytic role of the condensation domain. J. Biol. Chem. 273, 22773–22781 (1998).
Tseng, C. C. et al. Characterization of the surfactin synthetase C-terminal thioesterase domain as a cyclic depsipeptide synthase. Biochemistry 41, 13350–13359 (2002).
Krätzschmar, J., Krause, M. & Marahiel, M. A. Gramicidin S biosynthesis operon containing the structural genes grsA and grsB has an open reading frame encoding a protein homologous to fatty acid thioesterases. J. Bacteriol. 171, 5422–5429 (1989).
Hahn, M. & Stachelhaus, T. Harnessing the potential of communication-mediating domains for the biocombinatorial synthesis of nonribosomal peptides. Proc. Natl Acad. Sci. USA 103, 275–280 (2006).
Hoyer, K. M., Mahlert, C. & Marahiel, M. A. The iterative gramicidin S thioesterase catalyses peptide ligation and cyclization. Chem. Biol. 14, 13–22 (2007).
Gruenewald, S., Mootz, H. D., Stehmeier, P. & Stachelhaus, T. In vivo production of artificial nonribosomal peptide products in the heterologous host Escherichia coli. Appl. Environ. Microbiol. 70, 3282–3291 (2004).
Mootz, H. D. et al. Decreasing the ring size of a cyclic nonribosomal peptide antibiotic by in-frame module deletion in the biosynthetic genes. J. Am. Chem. Soc. 124, 10980–10981 (2002).
Nguyen, K. T. et al. Combinatorial biosynthesis of novel antibiotics related to daptomycin. Proc. Natl Acad. Sci. USA 103, 17462–17467 (2006).
Fischbach, M. A., Lai, J. R., Roche, E. D., Walsh, C. T. & Liu, D. R. Directed evolution can rapidly improve the activity of chimeric assembly-line enzymes. Proc. Natl Acad. Sci. USA 104, 11951–11956 (2007).
Butz, D. et al. Module extension of a non-ribosomal peptide synthetase of the glycopeptide antibiotic balhimycin produced by Amycolaptosis balhimycina. ChemBioChem 9, 1195–1200 (2008).
Calcott, M. J., Owen, J. G., Lamont, I. L. & Ackerley, D. F. Biosynthesis of novel pyoverdines by domain substitution in a nonribosomal peptide synthetase of Pseudomonas aeruginosa. Appl. Environ. Microbiol. 80, 5723–5731 (2014).
Murphy, A. C. Metabolic engineering is key to a sustainable chemical industry. Nat. Prod. Rep. 28, 1406–1425 (2011).
McKay, C. S. & Finn, M. G. Click chemistry in complex mixtures: bioorthogonal bioconjugation. Chem. Biol. 21, 1075–1101 (2014).
Jäckel, C. & Hilvert, D. Biocatalysts by evolution. Curr. Opin. Biotechnol. 21, 753–759 (2010).
Röttig, M. et al. NRPSpredictor2—a web server for predicting NRPS adenylation domain specificity. Nucleic Acids Res. 39, W362–W367 (2011).
Jin, M., Fischbach, M. A. & Clardy, J. A biosynthetic gene cluster for the acetyl-CoA carboxylase inhibitor andrimid. J. Am. Chem. Soc. 128, 10660–10661 (2006).
Acknowledgements
The authors thank A. Schütz and K. Malgorzata from the ETH Zurich Flow Cytometry Core Facility, C. Stutz-Ducommun and B. Blattmann from the Protein Crystallization Core Facility at the University of Zurich, N. Trapp from the Small Molecule Crystallography Center ETH Zurich, L. Bertschi from the Mass Spectrometry Service of the Laboratory of Organic Chemistry at ETH Zurich and the staff at the Swiss Light Source (Paul Scherrer Institute) for technical support. The authors also thank P. Mittl for assistance with data acquisition for X-ray crystallography, H.-M. Fischer for assistance with radiochemical experiments and J. Piel, P. Kast, T. Edwardson, X. Garrabou, S. Mantri and S. Studer for discussions. This work was supported by the ETH Zurich. Fellowships from the ETH (to D.A.H.), the Daiichi Sankyo Foundation of Life Science (to T.M.), the Scholarship Fund of the Swiss Chemical Industry (to H.K.) and the Studienstiftung des deutschen Volkes (to H.K.) are acknowledged.
Author information
Authors and Affiliations
Contributions
All authors were involved in project design. D.L.N., D.A.H., T.M., D.F. and H.K. executed experiments. The manuscript was written by D.L.N., D.A.H. and D.H., and revised by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 2541 kb)
Supplementary information
Crystallographic data for compound 9. (CIF 1683 kb)
Supplementary information
Crystallographic data for compound 10. (CIF 2692 kb)
Rights and permissions
About this article
Cite this article
Niquille, D., Hansen, D., Mori, T. et al. Nonribosomal biosynthesis of backbone-modified peptides. Nature Chem 10, 282–287 (2018). https://doi.org/10.1038/nchem.2891
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2891
This article is cited by
-
High-throughput engineering of biosynthetic assembly lines
Nature Chemical Biology (2024)
-
Adding α,α-disubstituted and β-linked monomers to the genetic code of an organism
Nature (2024)
-
High-throughput reprogramming of an NRPS condensation domain
Nature Chemical Biology (2024)
-
Genetically programmed cell-based synthesis of non-natural peptide and depsipeptide macrocycles
Nature Chemistry (2023)
-
Resurrecting ancestral antibiotics: unveiling the origins of modern lipid II targeting glycopeptides
Nature Communications (2023)