Abstract
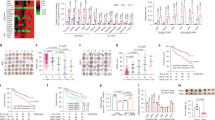
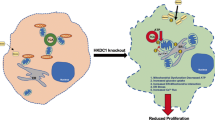
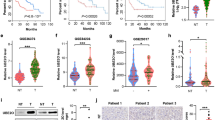
Dietary fructose is primarily metabolized in the liver. Here we demonstrate that, compared with normal hepatocytes, hepatocellular carcinoma (HCC) cells markedly reduce the rate of fructose metabolism and the level of reactive oxygen species, as a result of a c-Myc-dependent and heterogeneous nuclear ribonucleoprotein (hnRNP) H1- and H2-mediated switch from expression of the high-activity fructokinase (KHK)-C to the low-activity KHK-A isoform. Importantly, KHK-A acts as a protein kinase, phosphorylating and activating phosphoribosyl pyrophosphate synthetase 1 (PRPS1) to promote pentose phosphate pathway-dependent de novo nucleic acid synthesis and HCC formation. Furthermore, c-Myc, hnRNPH1/2 and KHK-A expression levels and PRPS1 Thr225 phosphorylation levels correlate with each other in HCC specimens and are associated with poor prognosis for HCC. These findings reveal a pivotal mechanism underlying the distinct fructose metabolism between HCC cells and normal hepatocytes and highlight the instrumental role of KHK-A protein kinase activity in promoting de novo nucleic acid synthesis and HCC development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Change history
27 April 2016
In the original version of this Article, the affiliations for Gilbert Cote, Hongxia Wang and Liwei Wang were incorrect. In addition, in the Acknowledgements section, 'National Science Foundation of China' should have read 'National Natural Science Foundation of China'. These errors have been corrected in all online versions.
References
Havel, P. J. Dietary fructose: implications for dysregulation of energy homeostasis and lipid/carbohydrate metabolism. Nutr. Rev. 63, 133–157 (2005).
Tappy, L. & Le, K. A. Metabolic effects of fructose and the worldwide increase in obesity. Physiol. Rev. 90, 23–46 (2010).
Lyssiotis, C. A. & Cantley, L. C. Metabolic syndrome: F stands for fructose and fat. Nature 502, 181–182 (2013).
Ishimoto, T. et al. Opposing effects of fructokinase C and A isoforms on fructose-induced metabolic syndrome in mice. Proc. Natl Acad. Sci. USA 109, 4320–4325 (2012).
Fong, Z. V. & Tanabe, K. K. The clinical management of hepatocellular carcinoma in the United States, Europe, and Asia: a comprehensive and evidence-based comparison and review. Cancer 120, 2824–2838 (2014).
Chan, S. L. et al. International validation of the Chinese university prognostic index for staging of hepatocellular carcinoma: a joint United Kingdom and Hong Kong study. Chin. J. Cancer 33, 481–491 (2014).
Zuo, T. T., Zheng, R. S., Zhang, S. W., Zeng, H. M. & Chen, W. Q. Incidence and mortality of liver cancer in China in 2011. Chin. J. Cancer 34, 56 (2015).
Bonthron, D. T., Brady, N., Donaldson, I. A. & Steinmann, B. Molecular basis of essential fructosuria: molecular cloning and mutational analysis of human ketohexokinase (fructokinase). Hum. Mol. Genet. 3, 1627–1631 (1994).
Hayward, B. E. & Bonthron, D. T. Structure and alternative splicing of the ketohexokinase gene. Eur. J. Biochem. 257, 85–91 (1998).
Diggle, C. P. et al. Ketohexokinase: expression and localization of the principal fructose-metabolizing enzyme. J. Histochem. Cytochem. 57, 763–774 (2009).
Asipu, A., Hayward, B. E., O’Reilly, J. & Bonthron, D. T. Properties of normal and mutant recombinant human ketohexokinases and implications for the pathogenesis of essential fructosuria. Diabetes 52, 2426–2432 (2003).
Lanaspa, M. A. et al. Endogenous fructose production and metabolism in the liver contributes to the development of metabolic syndrome. Nat. Commun. 4, 2434 (2013).
Ishimoto, T. et al. High-fat and high-sucrose (western) diet induces steatohepatitis that is dependent on fructokinase. Hepatology 58, 1632–1643 (2013).
Mirtschink, P. et al. HIF-driven SF3B1 induces KHK-C to enforce fructolysis and heart disease. Nature 522, 444–449 (2015).
Yang, W. et al. ERK1/2-dependent phosphorylation and nuclear translocation of PKM2 promotes the Warburg effect. Nat. Cell Biol. 14, 1295–1304 (2012).
Yang, W. & Lu, Z. Nuclear PKM2 regulates the Warburg effect. Cell Cycle 12, 3154–3158 (2013).
Kornberg, A., Lieberman, I. & Simms, E. S. Enzymatic synthesis and properties of 5-phosphoribosylpyrophosphate. J. Biol. Chem. 215, 389–402 (1955).
Khorana, H. G., Fernandes, J. F. & Kornberg, A. Pyrophosphorylation of ribose 5-phosphate in the enzymatic synthesis of 5-phosphorylribose 1-pyrophosphate. J. Biol. Chem. 230, 941–948 (1958).
Hove-Jensen, B. Mutation in the phosphoribosylpyrophosphate synthetase gene (prs) that results in simultaneous requirements for purine and pyrimidine nucleosides, nicotinamide nucleotide, histidine, and tryptophan in Escherichia coli. J. Bacteriol. 170, 1148–1152 (1988).
Nosal, J. M., Switzer, R. L. & Becker, M. A. Overexpression, purification, and characterization of recombinant human 5-phosphoribosyl-1-pyrophosphate synthetase isozymes I and II. J. Biol. Chem. 268, 10168–10175 (1993).
Li, S., Lu, Y., Peng, B. & Ding, J. Crystal structure of human phosphoribosylpyrophosphate synthetase 1 reveals a novel allosteric site. Biochem. J. 401, 39–47 (2007).
Li, B. et al. Negative feedback-defective PRPS1 mutants drive thiopurine resistance in relapsed childhood ALL. Nat. Med. 21, 563–571 (2015).
Cunningham, J. T., Moreno, M. V., Lodi, A., Ronen, S. M. & Ruggero, D. Protein and nucleotide biosynthesis are coupled by a single rate-limiting enzyme, PRPS2, to drive cancer. Cell 157, 1088–1103 (2014).
Mannava, S. et al. Direct role of nucleotide metabolism in C-MYC-dependent proliferation of melanoma cells. Cell Cycle 7, 2392–2400 (2008).
McMahon, S. B. Control of nucleotide biosynthesis by the MYC oncoprotein. Cell Cycle 7, 2275–2276 (2008).
Martinez-Contreras, R. et al. hnRNP proteins and splicing control. Adv. Exp. Med. Biol. 623, 123–147 (2007).
Keller, W. The RNA lariat: a new ring to the splicing of mRNA precursors. Cell 39, 423–425 (1984).
Rauch, J. et al. c-Myc regulates RNA splicing of the A-Raf kinase and its activation of the ERK pathway. Cancer Res. 71, 4664–4674 (2011).
Zimonjic, D. B. & Popescu, N. C. Role of DLC1 tumor suppressor gene and MYC oncogene in pathogenesis of human hepatocellular carcinoma: potential prospects for combined targeted therapeutics (review). Int. J. Oncol. 41, 393–406 (2012).
Trinh, C. H., Asipu, A., Bonthron, D. T. & Phillips, S. E. Structures of alternatively spliced isoforms of human ketohexokinase. Acta Crystallogr. D 65, 201–211 (2009).
Eriksen, T. A., Kadziola, A., Bentsen, A. K., Harlow, K. W. & Larsen, S. Structural basis for the function of Bacillus subtilis phosphoribosyl-pyrophosphate synthetase. Nat. Struct. Biol. 7, 303–308 (2000).
Willemoes, M., Hove-Jensen, B. & Larsen, S. Steady state kinetic model for the binding of substrates and allosteric effectors to Escherichia coli phosphoribosyl-diphosphate synthase. J. Biol. Chem. 275, 35408–35412 (2000).
Lim, J. S., Mietus-Snyder, M., Valente, A., Schwarz, J. M. & Lustig, R. H. The role of fructose in the pathogenesis of NAFLD and the metabolic syndrome. Nat. Rev. Gastroenterol. Hepatol. 7, 251–264 (2010).
Bose, T. & Chakraborti, A. S. Fructose-induced structural and functional modifications of hemoglobin: implication for oxidative stress in diabetes mellitus. Biochim. Biophys. Acta 1780, 800–808 (2008).
Bunn, H. F. & Higgins, P. J. Reaction of monosaccharides with proteins: possible evolutionary significance. Science 213, 222–224 (1981).
Wang, Y. et al. Prognostic significance of c-myc and AIB1 amplification in hepatocellular carcinoma. A broad survey using high-throughput tissue microarray. Cancer 95, 2346–2352 (2002).
Ji, H. et al. EGF-induced ERK activation promotes CK2-mediated disassociation of α-Catenin from β-Catenin and transactivation of β-Catenin. Mol. Cell 36, 547–559 (2009).
Xia, Y. et al. c-Jun downregulation by HDAC3-dependent transcriptional repression promotes osmotic stress-induced cell apoptosis. Mol. Cell 25, 219–232 (2007).
Dyballa, N. & Metzger, S. Fast and sensitive colloidal Coomassie G-250 staining for proteins in polyacrylamide gels. J. Vis. Exp. 30, 1431 (2009).
Lu, Z. et al. Activation of protein kinase C triggers its ubiquitination and degradation. Mol. Cell. Biol. 18, 839–845 (1998).
Kashima, T., Rao, N., David, C. J. & Manley, J. L. hnRNP A1 functions with specificity in repression of SMN2 exon 7 splicing. Hum. Mol. Genet. 16, 3149–3159 (2007).
Xu, Y. et al. Cell type-restricted activity of hnRNPM promotes breast cancer metastasis via regulating alternative splicing. Genes Dev. 28, 1191–1203 (2014).
Taus, T. et al. Universal and confident phosphorylation site localization using phosphoRS. J. Proteome Res. 10, 5354–5362 (2011).
Chen, Z. Y. et al. Dovitinib preferentially targets endothelial cells rather than cancer cells for the inhibition of hepatocellular carcinoma growth and metastasis. J. Transl. Med. 10, 245 (2012).
Yang, W. et al. Nuclear PKM2 regulates β-catenin transactivation upon EGFR activation. Nature 480, 118–122 (2011).
Acknowledgements
We thank B.-F. Pan for technical support and D. Norwood for critical reading of this manuscript. This work was supported by National Cancer Institute grants 2R01 CA109035 (Z.L.) and 1R0 CA169603 (Z.L.), National Institute of Neurological Disorders and Stroke grant 1R01 NS089754 (Z.L.), MD Anderson Support Grant CA016672, the James S. McDonnell Foundation 21st Century Science Initiative in Brain Cancer Research Award 220020318 (Z.L.), 2P50 CA127001 (Brain Cancer SPORE), a Sister Institution Network Fund from MD Anderson (Z.L.; C.-N.Q.), NIH High-End Instrumentation program grant 1S10OD012304-01 (D.H.H.), CPRIT Core Facility Grant RP130397 (D.H.H.), the Odyssey Fellowship from MD Anderson (X.L.), and research grants 81272340 (C.-N.Q.) and 81472386 (C.-N.Q.) from the National Natural Science Foundation of China. Z.L. is a Ruby E. Rutherford Distinguished Professor.
Author information
Authors and Affiliations
Contributions
This study was conceived by Z.L. and X.L.; Z.L. and X.L. designed the study; X.L., X.Q., L.-X.P., Y.J., D.H.H., Y.Z., Y.X., J.-H.L. and G.C. performed experiments; L.W., H.W. and C.-N.Q. provided reagents and technical assistance; Z.L. and X.L. wrote the paper with comments from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 5 hnRNPH1/2 expression switches KHK-C expression to KHK-A expression in HCC cells.
(a) Total RNA was extracted from 30 human HCC and paired non-tumor tissue samples. The RT-PCR products were digested by BsmAI and resolved on agarose gels. N, non-tumor tissue; T, tumor tissue. (b) Immunoprecipitation of hnRNPH1/2 (bottom left panel) from Hep3B cells was followed by RT-PCR analysis of hnRNPH1/2-associated RNAs (bottom right panel) with the indicated primers (top panel). (c) Amino acid sequences of hnRNPH1 and hnRNPH2 were aligned. Different amino acids in two proteins are highlighted. (d) Schematic diagram of synthesized pre-mRNAs of AdML and the indicated AdML-KHK exon 3C fused products (left panel). Incubation of the synthesized pre-mRNAs with a Hep3B nuclear extract with or without hnRNPH1/2 depletion was followed by PCR analysis (right panel).
Supplementary Figure 6 KHK-A but not KHK-C phosphorylates PRPS1 at Thr225.
(a) Purified recombinant GST-KHK-A was incubated with immobilized purified recombinant His-PRPS1 or His-PRPS2. An in vitro pulldown assay was performed. Immunoblot analysis was conducted with the indicated antibodies. (b) Schematic diagram of the restriction site of HindIII in PRPS1 cDNA (left panel). Total RNA was extracted from normal hepatocytes and the indicated HCC cells (right panel). (c) The Km of purified GST-KHK-A for fructose with representative plotting of 1/V vs. 1/[fructose] (left panel) or for purified His-PRPS1 with representative plotting of 1/CPM vs. 1/[PRPS1] (right panel) was determined. (d) Hep3 cells were centrifuged and washed with PBS. 100 μl of the cell pellets with removal of PBS were mixed with 900 μl lysate buffer. 10 μl cell lysate was mixed with 10 μl of purified recombinant his-PRPS1 with the indicated concentrations were analyzed by immunoblot assay with an anti-PRPS antibody. (e) Bacterially purified indicated KHK-A and PRPS1 proteins were stained with colloidal coomassie blue. (f) In vitro phosphorylation analysis, SDS-PAGE and autoradiography were performed by mixing purified GST-KHK-A with purified His-PRPS1 or His-PRPS2 in the presence of [γ32P]-ATP. Immunoblot analysis was conducted with the indicated antibodies. (g) Hep3B cells were transiently transfected with a vector expressing the indicated Flag-PRPS proteins. Immunoblot analysis was conducted with the indicated antibodies. (h) GST pull-down analysis was performed by mixing purified immobilized WT GST-KHK-A, GST-KHK-A G257R on glutathione agarose beads with purified His-PRPS1. (i) [γ32P]-ATP was mixed with WT KHK-A or the KHK-A G257R mutant. KHK-A–bound [γ32P]-ATP was measured. The data represent the mean ± SD from n = 3 independent experiments. (j) In vitro phosphorylation assays were performed by mixing purified WT KHK-A or KHK-C with purified PRPS1 in the presence of ATP. Immunoblot analysis was conducted with the indicated antibodies (k) Hep3B cells were cultured in the medium with added fructose with the indicated concentrations. Immunoblot analysis was conducted with the indicated antibodies.
Supplementary Figure 7 Phosphorylated PRPS1 was not regulated by the allosteric regulators.
(a) Activity of bacterially purified WT His-PRPS1 was measured in the presence or absence of phosphate (50 mM) ADP (100 μM). The data represent the mean ± SD from n = 3 independent experiments. (b) The activity of bacterially purified WT His-PRPS1 with or without phosphorylation by GST-KHK-A was measured in the presence or absence of IMP (100 μM), AMP (100 μM), or GMP (100 μM). The data represent the mean ± SD from n = 3 independent experiments. Immunoblot analysis was conducted with the indicated antibodies. (c) The activity of bacterially purified WT His-PRPS1 was measured in the presence or absence of phosphate (50 mM), IMP (100 μM), AMP (100 μM), or GMP (100 μM). The data represent the mean ± SD from n = 3 independent experiments. Immunoblot analysis was conducted with the indicated antibody. (d) Immunoprecipitation analyses with PRPS1 pT225 antibody was performed using Hep3B cell lysates. Immunoblot analysis was conducted with the indicated antibodies. The amount of phosphorylated PRPS1 was quantified. HC represents heavy chain.
Supplementary Figure 8 KHK-A-mediated PRPS1 phosphorylation promotes glucose-derived de novo nucleotide synthesis.
(a) Intracellular IMP levels in lysates of Hep3B cells expressing hnRNPH1/2 shRNA were measured. The data represent the mean ± SD from n = 3 independent experiments. A two-tailed Student’s t test was used. ∗ represents P < 0.01 between the cells with or without hnRNPH1/2 depletion. (b) Intracellular IMP levels in lysates of Hep3B cells expressing KHK shRNA with or without reconstituted expression of WT rKHK-A, rKHK-A L83A, or WT rKHK-C were measured. The data represent the mean ± SD from n = 3 independent experiments. A two-tailed Student’s t test was used. ∗ represents P < 0.01 between the cells with or without KHK depletion. #represents P < 0.01 between the KHK-depleted cells reconstituted expression of WT rKHK-A and the KHK-depleted cells reconstituted expression of rKHK-A L83A and WT rKHK-C. (c) Intracellular IMP levels in lysates of Hep3B cells expressing PRPS1 shRNA with or without reconstituted expression of WT rPRPS1, rPRPS1 T225A, rPRPS1 A190T were measured. The data represent the mean ± SD from n = 3 independent experiments. A two-tailed Student’s t test was used. ∗ represents P < 0.01 between the cells with or without PRPS1 depletion. #represents P < 0.01 between the PRPS1-depleted cells reconstituted expression of WT rPRPS1 and the rPRPS1 T225A. (d,e) Hep3B cells with reconstituted expression of KHK-C or KHK-A were cultured in the presence or absence of fructose (10 mM). The intracellular levels of AT (d) and phosphate (e) were measured. The data represent the mean ± SD from n = 3 independent experiments. A two-tailed Student’s t test was used. ∗ represents P < 0.01; NS represents not significant difference between the indicated cells with or without fructose treatment.
Supplementary Figure 9 KHK-A-mediated PRPS1 phosphorylation promotes hepatocellular tumorigenesis and is associated with the pathogenesis of HCC.
(a) The apoptosis of Hep3B and Huh-7 cells with depletion of hnRNP1/2 (left panel), KHK-A (middle panel), and PRPS1 (right panel) and with or without reconstituted expression of the indicated proteins were examined. The data represent the mean ± SD from n = 3 independent experiments. (b) The proliferation rates of Hep3B cells with the depletion of KHK-A and reconstituted expression of the indicated proteins were examined in the presence or absence of added fructose (10 mM) and NAC (2 mM). The data represent the mean ± SD from n = 3 independent experiments. ∗ P < 0.05. (c) Lysates of tumors derived from Huh-7 cells with KHK or PRPS1 depletion and with or without reconstituted WT rKHK-A, rKHK-A L83A, WT rKHK-C, WT rPRPS1, or rPRPS1 T225A expression were prepared. Immunoblot analyses with the indicated antibodies were conducted (d) The antibody specificities were validated. Immunohistochemical analysis of the tissues from the tumors derived from Huh-7 cells (left and right panel) or Huh-7 cells with KHK depletion and reconstituted expression of WT rKHK-C (middle panel) with the indicated antibodies was performed in the presence or absence of blocking peptides specific for KHK Exon 3A- or Exon 3C-coded regions or phosphorylated PRPS1 T225. (e) IHC staining of n = 90 human HCC samples with anti-phospho-PRPS1 T225, anti-KHK-A, anti-hnRNPH1/2, and anti-c-Myc antibodies was performed. Chi-square analysis was performed depending on the staining score for each antibody (high staining score 4.1-8.0; low staining score 0-4).
Supplementary information
Supplementary Information
Supplementary Information (PDF 2990 kb)
Rights and permissions
About this article
Cite this article
Li, X., Qian, X., Peng, LX. et al. A splicing switch from ketohexokinase-C to ketohexokinase-A drives hepatocellular carcinoma formation. Nat Cell Biol 18, 561–571 (2016). https://doi.org/10.1038/ncb3338
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3338
This article is cited by
-
The polyol pathway and nuclear ketohexokinase A signaling drive hyperglycemia-induced metastasis of gastric cancer
Experimental & Molecular Medicine (2024)
-
Direct stimulation of de novo nucleotide synthesis by O-GlcNAcylation
Nature Chemical Biology (2024)
-
Rare sugar l-sorbose exerts antitumor activity by impairing glucose metabolism
Communications Biology (2023)
-
Nucleus-exported CLOCK acetylates PRPS to promote de novo nucleotide synthesis and liver tumour growth
Nature Cell Biology (2023)
-
MYC in liver cancer: mechanisms and targeted therapy opportunities
Oncogene (2023)