Abstract
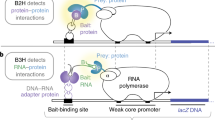
We have modified the yeast two-hybrid system to enable the detection of protein-protein interactions that require a specific post-translational modification, using the acetylation of histones and the phosphorylation of the carboxyl terminal domain (CTD) of RNA polymerase II as test modifications. In this tethered catalysis assay, constitutive modification of the protein to be screened for interactions is achieved by fusing it to its cognate modifying enzyme, with the physical linkage resulting in efficient catalysis. This catalysis maintains substrate modification even in the presence of antagonizing enzyme activities. A catalytically inactive mutant of the enzyme is fused to the substrate as a control such that the modification does not occur; this construct enables the rapid identification of modification-independent interactions. We identified proteins with links to chromatin functions that interact with acetylated histones, and proteins that participate in RNA polymerase II functions and in CTD phosphorylation regulation that interact preferentially with the phosphorylated CTD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Tucker, C.L., Gera, J.F. & Uetz, P. Towards an understanding of complex protein networks. Trends Cell Biol. 11, 102–106 (2001).
Yaffe, M.B. How do 14-3-3 proteins work? Gatekeeper phosphorylation and the molecular anvil hypothesis. FEBS Lett. 513, 53–57 (2002).
Cattaneo, E. & Pelicci, P.G. Emerging roles for SH2/PTB-containing Shc adaptor proteins in the developing mammalian brain. Trends Neurosci. 21, 476–481 (1998).
Fischle, W., Wang, Y. & Allis, C.D. Histone and chromatin cross-talk. Curr. Opin. Cell Biol. 15, 172–183. (2003).
Carmen, A.A., Milne, L. & Grunstein, M. Acetylation of the yeast histone H4 N terminus regulates its binding to heterochromatin protein SIR3. J. Biol. Chem. 277, 4778–4781 (2002).
Fields, S. & Song, O. A novel genetic system to detect protein-protein interactions. Nature 340, 245–246 (1989).
Osborne, M.A., Dalton, S. & Kochan, J.P. The yeast tribrid system—genetic detection of trans-phosphorylated ITAM-SH2-interactions. Biotechnology (N Y) 13, 1474–1478 (1995).
Shaywitz, A.J., Dove, S.L., Kornhauser, J.M., Hochschild, A. & Greenberg, M.E. Magnitude of the CREB-dependent transcriptional response is determined by the strength of the interaction between the kinase-inducible domain of CREB and the KIX domain of CREB-binding protein. Mol. Cell Biol. 20, 9409–9422 (2000).
Yamada, M. et al. Analysis of tyrosine phosphorylation-dependent protein-protein interactions in TrkB-mediated intracellular signaling using modified yeast two-hybrid system. J. Biochem. (Tokyo) 130, 157–165 (2001).
Kornbluth, S., Jove, R. & Hanafusa, H. Characterization of avian and viral p60src proteins expressed in yeast. Proc. Natl. Acad. Sci. USA 84, 4455–4459 (1987).
Hardwick, K.G., Weiss, E., Luca, F.C., Winey, M. & Murray, A.W. Activation of the budding yeast spindle assembly checkpoint without mitotic spindle disruption. Science 273, 953–956 (1996).
Xu, P., Jacobs, A.R. & Taylor, S.I. Interaction of insulin receptor substrate 3 with insulin receptor, insulin receptor-related receptor, insulin-like growth factor-1 receptor, and downstream signaling proteins. J. Biol. Chem. 274, 15262–15270 (1999).
Candau, R., Zhou, J.X., Allis, C.D. & Berger, S.L. Histone acetyltransferase activity and interaction with ADA2 are critical for GCN5 function in vivo. EMBO J. 16, 555–565 (1997).
Kuo, M.H., Zhou, J., Jambeck, P., Churchill, M.E. & Allis, C.D. Histone acetyltransferase activity of yeast Gcn5p is required for the activation of target genes in vivo. Genes Dev. 12, 627–639 (1998).
Sterner, D.E. & Berger, S.L. Acetylation of histones and transcription-related factors. Microbiol. Mol. Biol. Rev. 64, 435–459 (2000).
Broder, Y.C., Katz, S. & Aronheim, A. The ras recruitment system, a novel approach to the study of protein-protein interactions. Curr. Biol. 8, 1121–1124 (1998).
James, P., Halladay, J. & Craig, E.A. Genomic libraries and a host strain designed for highly efficient two-hybrid selection in yeast. Genetics 144, 1425–1436 (1996).
Kassenbrock, C.K. et al. RPM2, independently of its mitochondrial RNase P function, suppresses an ISP42 mutant defective in mitochondrial import and is essential for normal growth. Mol. Cell. Biol. 15, 4763–4770 (1995).
Ho, Y. et al. Systematic identification of protein complexes in Saccharomyces cerevisiae by mass spectrometry. Nature 415, 180–183 (2002).
Uetz, P. et al. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000).
Marcus, G.A., Silverman, N., Berger, S.L., Horiuchi, J. & Guarente, L. Functional similarity and physical association between GCN5 and ADA2: putative transcriptional adaptors. EMBO J. 13, 4807–4815 (1994).
Recht, J., Dunn, B., Raff, A. & Osley, M.A. Functional analysis of histones H2A and H2B in transcriptional repression in Saccharomyces cerevisiae. Mol. Cell Biol. 16, 2545–2553 (1996).
Georgel, P.T., Tsukiyama, T. & Wu, C. Role of histone tails in nucleosome remodeling by Drosophila NURF. EMBO J. 16, 4717–4726 (1997).
Shen, E.C. et al. Arginine methylation facilitates the nuclear export of hnRNP proteins. Genes Dev. 12, 679–691 (1998).
Lacoste, N., Utley, R.T., Hunter, J.M., Poirier, G.G. & Cote, J. Disruptor of Telomeric Silencing-1 Is a Chromatin-specific Histone H3 Methyltransferase. J. Biol. Chem. 277, 30421–30424 (2002).
Ladurner, A.G., Inouye, C., Jain, R. & Tjian, R. Bromodomains mediate an acetyl-histone encoded antisilencing function at heterochromatin boundaries. Mol. Cell 11, 365–376 (2003).
Kobor, M.S. & Greenblatt, J. Regulation of transcription elongation by phosphorylation. Biochim. Biophys. Acta 1577, 261–275 (2002).
Kobor, M.S. et al. An unusual eukaryotic protein phosphatase required for transcription by RNA polymerase II and CTD dephosphorylation in S. cerevisiae. Mol. Cell 4, 55–62 (1999).
Valay, J.G., Simon, M. & Faye, G. The kin28 protein kinase is associated with a cyclin in Saccharomyces cerevisiae. J. Mol. Biol. 234, 307–310 (1993).
Sherman, F. Getting started with yeast. Methods Enzymol. 194, 3–21 (1991).
Acknowledgements
We thank David Arnosti and Steven Triezenberg for critical reading of this manuscript, Steven Triezenberg for suggesting the name of 'tethered catalysis,' Asha Acharya for chicken histones and bacterial lysate preparation, Kanchan Champhekar for CTD-Kin28 plasmid construction and members of the Triezenberg and Kuo labs for helpful discussion throughout this work. This work was supported by National Institutes of Health grant R01 GM62282 (M.-H.K.) and P41 RR11823 (S.F.). S.F. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Summary of acetylated histone-binding proteins identified by the array approach. (PDF 180 kb)
Supplementary Table 2
Summary of phosphorylated CTD-binding proteins identified by the array approach. (PDF 104 kb)
Supplementary Figure 1
Growth tests of various yeast ORFs for their abilities to interact with CTD or Kin28. (PDF 501 kb)
Rights and permissions
About this article
Cite this article
Guo, D., Hazbun, T., Xu, XJ. et al. A tethered catalysis, two-hybrid system to identify protein-protein interactions requiring post-translational modifications. Nat Biotechnol 22, 888–892 (2004). https://doi.org/10.1038/nbt985
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt985
This article is cited by
-
Post-translational selective intracellular silencing of acetylated proteins with de novo selected intrabodies
Nature Methods (2017)
-
The regulatory role of Pcf11-similar-4 (PCFS4) in Arabidopsis development by genome-wide physical interactions with target loci
BMC Genomics (2013)
-
Identification of cellular factors binding to acetylated HIV-1 integrase
Amino Acids (2011)
-
GCN5-dependent acetylation of HIV-1 integrase enhances viral integration
Retrovirology (2010)
-
Systems for the detection and analysis of protein–protein interactions
Applied Microbiology and Biotechnology (2006)