Abstract
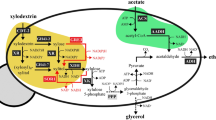
We obtained efficient conversion of xylose to xylitol by transforming Saccharomyces cerevisiae with the gene encoding the xylose reductase (XR) of Pichia stipitis CBS 6054. Comparison of the chromosomal and cDNA copies of the XYL1 gene showed that the genomic XYL1 contains no introns, and an XR monomer of 318 amino acids (35,985 D) is encoded by an open reading frame of 954 bp. The amino acid sequence of the P. stipitis XR is similar to several aldose reductases, suggesting that P. stipitis XR is part of the aldoketo reductase superfamily. S. cerevisiae transformed with the XYL1 gene gave over 95% conversion of xylose into xylitol, a yield not obtainable with natural xylose utilizing yeasts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Hyvönen, L. and Koivistoinen, P. 1983. Food technological evaluation of xylitol. Adv. in Food Research 28: 373–403.
Melaja, A. and Hämäläinen, L. 1977. Process for making xylitol. US Patent 4,008,285.
Jeffries, T.W. 1990. Fermentation of D-xylose and cellobiose. p. 349–394 In: Yeast: Biotechnology and Biocatalysis. Verachtert, H. and De Mot, R. (Eds.). Marcel Dekker, Inc., NY.
Verduyn, C., Frank, J., Jzn Van Dijken, J.P. and Scheffers, W.A. 1985. Multiple forms of xylose reductase in Pachysolen tannophilus. FEMS Microbiol. Lett. 30: 313–317.
Bruinenberg, P.M., de Bot, P.M., van Dijken, J.P. and Scheffers, W.A. 1984. NADH-linked aldose reductase: The key to anaerobic alcoholic fermentation of xylose by yeasts. Appl. Microbiol. Biotechnol. 19: 256–260.
Ojamo, H., Ylinen, L. and Linko, M. 1987. Mikrobiologinen valmistusmenetelmä. Finnish patent 76377.
Meyrial, V., Delgenes, J.P., Moletta, R., Navarro, J.M. and Inra, I.A.A. 1991. Xylitol production from D-xylose by Candida guillermondii—Fermentation behaviour. Biotechnol. Lett. 13: 281–286.
Verduyn, C., Van Kleff, R., Schreuder, H., Van Dijken, J.P. and Scheffers, W.A. 1985. Properties of the NAD(P)H-dependent xylose reductase from the xylose fermenting yeast Pichia stipitis. Biochem. J. 226: 669–677.
Rizzi, M., Erlemann, P., Bui-Thanh, N.-A. and Dellweg, H. 1988. Xylose fermentation by yeasts 4. Purification and kinetic studies of xylose reductase from Pichia stipitis. Appl. Microbiol. Biotechnol. 29: 148–154.
Ditzelmüller, G., Kubicek, C.P., Wöhrer, W. and Röhr, M. 1984. Xylose metabolism in Pachysolen tannophilus: Purification and properties of xylose reductase. Can. J. Microbiol. 30: 1330–1336.
Sharp, P., Cowe, E., Higgins, D.G., Shields, D.C., Wolfe, K.H. and Wright, F. 1988. Codon usage patterns in Escherichia coli, Bacillus subtilis, Schizosaccharomyes pombe, Drosophila melanogaster and Homo sapiens; a review of the considerable within-species diversity. Nuc. Acids Res. 16: 8207–8211.
Bohren, K.M., Bullock, B., Wermuth, B. and Gabbay, K.M. 1989. The aldo-keto reductase superfamily cDNAs and deduced amino acid sequences of human aldehyde and aldose reductases. J. Biol. Chem. 264: 9547–9551.
Carper, D., Nishimura, C., Shinohara, T., Dietzchold, B., Wistow, G., Kador, P. and Kinoshita, I.H. 1987. Aldose reductase and rho-crystallin belong to the same protein superfamily as aldehyde reductase. FEES Lett. 220: 209–213.
Petrash, J.M. and Favello, A.D. 1989. Isolation and characterization of cDNA clones encoding aldose reductase. Curr. Eye Res. 8: 1021–1027.
Watanabe, K., Fuji, Y., Nakayama, K., Okhuba, H., Kuramitsu, S., Kagamiyama, H., Nakanishi, S. and Hayashi, O. 1988. Structural similarity of bovine lung prostaglandin F synthase to lens epsilon-crystallin of the European common frog. Proc. Natl. Acad. Sci. USA 85: 11–15.
Winters, C.J., Molowa, D.T. and Guzelian, P.S. 1990. Isolation and characterization of cloned cDNAs encoding human liver chlordecone reductase. Biochemistry 29: 1080–1087.
Ho, N.W.Y., Lin, F.P., Huang, S., Andrews, P.C. and Tsao, G.T. 1990. Purification, characterization, and amino acid terminal sequence of xylose reductase from Candida shehatae. Enzyme Microb. Technol. 12: 33–39.
Kurtzman, C.P. 1990. Candida shehatae—genetic diversity and phylo-genetic relationships with other xylose-fermenting yeasts. Antonie van Leeuwenhoek 57: 215–222.
Prior, B.A., Kilian, S.G. and du Perez, J.C. 1989. Fermentation of D-xylose by the yeasts Candida shehatae and Pichia stipitis. Process. Biochem. 24: 21–32.
Kötter, P., Amore, R., Hollenberg, C.P. and Ciriacy, M. 1990. Isolation and characterization of the Pichia stipitis xylitol dehydroge-nase gene, XYL2, and construction of a xylose-utilizing Saccharomyces cerevisiae transformant. Curr. Genet. 18: 493–500.
Van Zyl, C., Prior, O.L., Kilian, S.G. and Kock, J.L.F. 1989. D-Xylose utilization by Saccharomyces cerevisiae. J. Gen. Microbiol. 135: 2791–2798.
Barbosa, M.F.S., de Medeiros, M.B., de Manchila, I.M., Schneider, H. and Lee, H. 1988. Screening for yeasts for production of xylitol from D-xylose and some factors which affect xylitol yield in Candida guilliermondii. J. Ind. Microbiol. 3: 241–251.
Mellor, J., Dobson, M.J., Roberts, N.A., Tuite, M.F., Emtage, J.S., White, S., Lowe, P.A., Patel, T., Kingsman, A.J. and Kingsman, S.M. 1983. Efficient synthesis of enzymatically active calf chymosin in Saccharomyces cerevisiae. Gene 24: 1–14.
Bradford, M.M. 1976. A rapid and sensitive method for the quantification of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72: 248–254.
Smiley, K.L. and Bolen, P.L. 1982. Demonstration of D-xylose reductase and D-xylitol dehydrogenase in Pachysolen tannophilus. Biotechnol. Lett. 4: 607–610.
Laemmli, U.K. 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685.
Shaw, C.R. and Prasad, R. 1970. Starch electrophoresis of enzymes—a compilation of recipes. Biochem. Genet. 4: 297–320.
Chirgwin, J.M., Przybyla, A.E., MacDonald, R.J. and Rutter, W.J. 1979. Isolation of biologically active ribonucleic acid from sources enriched in ribonuclease. Biochemistry 18: 5294–5299.
Young, R.A. and Davis, R.W. 1983. Yeast RNA polymerase II genes: isolation with antibody probes. Science 222: 778–782.
Güssow, D. and Clackson, T. 1989. Direct clone characterization from plaques and colonies by the polymerase chain reaction. Nuc. Acid Res. 17: 4000.
Cryer, D.R., Eccleshall, R. and Marmur, J. 1975. Isolation of yeast DNA. Methods Cell Biol. 12: 39–44.
Zagursky, R.I., Berman, M.L., Baumeister, K. and Lomax, N. 1986. Rapid and easy sequencing of large linear double stranded DNA and supercoiled plasmid DNA. Gene Anal. Techn. 2: 89–94.
Devereux, J., Haberli, P. and Smithies, O. 1984. A comprehensive set of sequence analysis programs for the VAX.Nuc. Acids Res. 12: 387–395.
Ito, H., Fukuda, M., Murata, K. and Kimura, A. 1983. Transformation of intact yeast cells treated with alkali cations. J. Bact. 153: 163–168.
Sherman, F., Fink, G. and Hicks, I.B. 1983 Methods in Yeast Genetics. A Laboratory Manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hallborn, J., Walfridsson, M., Airaksinen, U. et al. Xylitol Production by Recombinant Saccharomyces Cerevisiae. Nat Biotechnol 9, 1090–1095 (1991). https://doi.org/10.1038/nbt1191-1090
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt1191-1090