Abstract
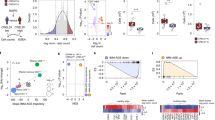
Natural IgM are involved in numerous immunological functions but the genetic factors that control the homeostasis of its secretion and upholding remain unknown. Prompted by the finding that C57BL/6 mice had significantly lower serum levels of IgM when compared with BALB/c mice, we performed a genome-wide screen and found that the level of serum IgM was controlled by a QTL on chromosome 13 reaching the highest level of association at marker D13Mit266 (LOD score=3.54). This locus was named IgMSC1 and covered a region encompassing the interferon-regulatory factor 4 gene (Irf4). The number of splenic mature B cells in C57BL/6 did not differ from BALB/c mice but we found that low serum levels of IgM in C57BL/6 mice correlated with lower frequency of IgM-secreting cells in the spleen and in the peritoneal cavity. These results suggested that C57BL/6 mice have lower efficiency in late B-cell maturation, a process that is highly impaired in Irf4 knockout mice. In fact, we also found reduced Irf4 gene expression in B cells of C57BL/6 mice. Thus, we propose Irf4 as a candidate for the IgMSC1 locus, which controls IgM homeostatic levels at the level of B-cell terminal differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Boes M, Prodeus AP, Schmidt T, Carroll MC, Chen J . A critical role of natural immunoglobulin M in immediate defense against systemic bacterial infection. J Exp Med 1998; 188: 2381–2386.
Brown JS, Hussell T, Gilliland SM, Holden DW, Paton JC, Ehrenstein MR et al. The classical pathway is the dominant complement pathway required for innate immunity to Streptococcus pneumoniae infection in mice. Proc Natl Acad Sci USA 2002; 99: 16969–16974.
Ochsenbein AF, Fehr T, Lutz C, Suter M, Brombacher F, Hengartner H et al. Control of early viral and bacterial distribution and disease by natural antibodies. Science 1999; 286: 2156–2159.
Boes M, Schmidt T, Linkemann K, Beaudette BC, Marshak-Rothstein A, Chen J . Accelerated development of IgG autoantibodies and autoimmune disease in the absence of secreted IgM. Proc Natl Acad Sci USA 2000; 97: 1184–1189.
Ehrenstein MR, Cook HT, Neuberger MS . Deficiency in serum immunoglobulin (Ig)M predisposes to development of IgG autoantibodies. J Exp Med 2000; 191: 1253–1258.
Boes M, Esau C, Fischer MB, Schmidt T, Carroll M, Chen J . Enhanced B-1 cell development, but impaired IgG antibody responses in mice deficient in secreted IgM. J Immunol 1998; 160: 4776–4787.
Ehrenstein MR, O'Keefe TL, Davies SL, Neuberger MS . Targeted gene disruption reveals a role for natural secretory IgM in the maturation of the primary immune response. Proc Natl Acad Sci USA 1998; 95: 10089–10093.
Baker N, Ehrenstein MR . Cutting edge: selection of B lymphocyte subsets is regulated by natural IgM. J Immunol 2002; 169: 6686–6690.
Brandlein S, Pohle T, Ruoff N, Wozniak E, Muller-Hermelink HK, Vollmers HP . Natural IgM antibodies and immunosurveillance mechanisms against epithelial cancer cells in humans. Cancer Res 2003; 63: 7995–8005.
Vollmers HP, Brandlein S . The ‘early birds’: natural IgM antibodies and immune surveillance. Histol Histopathol 2005; 20: 927–937.
Peng Y, Kowalewski R, Kim S, Elkon KB . The role of IgM antibodies in the recognition and clearance of apoptotic cells. Mol Immunol 2005; 42: 781–787.
Lacroix-Desmazes S, Kaveri SV, Mouthon L, Ayouba A, Malanchere E, Coutinho A et al. Self-reactive antibodies (natural autoantibodies) in healthy individuals. J Immunol Methods 1998; 216: 117–137.
Bos NA, Meeuwsen CG, Hooijkaas H, Benner R, Wostmann BS, Pleasants JR . Early development of Ig-secreting cells in young of germ-free BALB/c mice fed a chemically defined ultrafiltered diet. Cell Immunol 1987; 105: 235–245.
Haury M, Sundblad A, Grandien A, Barreau C, Coutinho A, Nobrega A . The repertoire of serum IgM in normal mice is largely independent of external antigenic contact. Eur J Immunol 1997; 27: 1557–1563.
Boes M . Role of natural and immune IgM antibodies in immune responses. Mol Immunol 2000; 37: 1141–1149.
Lydyard PM, Quartey-Papafio R, Broker B, Mackenzie L, Jouquan J, Blaschek MA et al. The antibody repertoire of early human B cells. I. High frequency of autoreactivity and polyreactivity. Scand J Immunol 1990; 31: 33–43.
Manson JJ, Mauri C, Ehrenstein MR . Natural serum IgM maintains immunological homeostasis and prevents autoimmunity. Springer Semin Immunopathol 2005; 26: 425–432.
Shapiro-Shelef M, Calame K . Regulation of plasma-cell development. Nat Rev Immunol 2005; 5: 230–242.
Srivastava B, Quinn III WJ, Hazard K, Erikson J, Allman D . Characterization of marginal zone B cell precursors. J Exp Med 2005; 202: 1225–1234.
Chung JB, Silverman M, Monroe JG . Transitional B cells: step by step towards immune competence. Trends Immunol 2003; 24: 343–349.
Martin F, Kearney JF . Marginal-zone B cells. Nat Rev Immunol 2002; 2: 323–335.
Martin F, Kearney JF . CD21high IgMhigh splenic B cells enriched in the marginal zone: distinct phenotypes and functions. Curr Top Microbiol Immunol 1999; 246: 45–50; discussion 1–2.
Oliver AM, Martin F, Gartland GL, Carter RH, Kearney JF . Marginal zone B cells exhibit unique activation, proliferative and immunoglobulin secretory responses. Eur J Immunol 1997; 27: 2366–2374.
Holmberg D, Freitas AA, Portnoi D, Jacquemart F, Avrameas S, Coutinho A . Antibody repertoires of normal BALB/c mice: B lymphocyte populations defined by state of activation. Immunol Rev 1986; 93: 147–169.
Pereira P, Forni L, Larsson EL, Cooper M, Heusser C, Coutinho A . Autonomous activation of B and T cells in antigen-free mice. Eur J Immunol 1986; 16: 685–688.
Herzenberg LA . B-1 cells: the lineage question revisited. Immunol Rev 2000; 175: 9–22.
Hardy RR, Hayakawa K . CD5 B cells, a fetal B cell lineage. Adv Immunol 1994; 55: 297–339.
Tumang JR, Frances R, Yeo SG, Rothstein TL . Spontaneously Ig-secreting B-1 cells violate the accepted paradigm for expression of differentiation-associated transcription factors. J Immunol 2005; 174: 3173–3177.
Calame KL, Lin KI, Tunyaplin C . Regulatory mechanisms that determine the development and function of plasma cells. Annu Rev Immunol 2003; 21: 205–230.
Klein U, Casola S, Cattoretti G, Shen Q, Lia M, Mo T et al. Transcription factor IRF4 controls plasma cell differentiation and class-switch recombination. Nat Immunol 2006; 7: 773–782.
Mamane Y, Heylbroeck C, Genin P, Algarte M, Servant MJ, LePage C et al. Interferon regulatory factors: the next generation. Gene 1999; 237: 1–14.
Sciammas R, Shaffer AL, Schatz JH, Zhao H, Staudt LM, Singh H . Graded expression of interferon regulatory factor-4 coordinates isotype switching with plasma cell differentiation. Immunity 2006; 25: 225–236.
Pernis AB . The role of IRF-4 in B and T cell activation and differentiation. J Interferon Cytokine Res 2002; 22: 111–120.
Marecki S, Fenton MJ . The role of IRF-4 in transcriptional regulation. J Interferon Cytokine Res 2002; 22: 121–133.
Mittrucker HW, Matsuyama T, Grossman A, Kundig TM, Potter J, Shahinian A et al. Requirement for the transcription factor LSIRF/IRF4 for mature B and T lymphocyte function. Science 1997; 275: 540–543.
Lin L, Gerth AJ, Peng SL . Active inhibition of plasma cell development in resting B cells by microphthalmia-associated transcription factor. J Exp Med 2004; 200: 115–122.
Grundbacher FJ . Heritability estimates and genetic and environmental correlations for the human immunoglobulins G, M, and A. Am J Hum Genet 1974; 26: 1–12.
Barbosa CA, Rao DC, Morton NE . Analysis of family resemblance for immunoglobulin M, G and A levels. Hum Hered 1981; 31: 8–14.
Borecki IB, McGue M, Gerrard JW, Lebowitz MD, Rao DC . Familial resemblance for immunoglobulin levels. Hum Genet 1994; 94: 179–185.
Barnes KC, Neely JD, Duffy DL, Freidhoff LR, Breazeale DR, Schou C et al. Linkage of asthma and total serum IgE concentration to markers on chromosome 12q: evidence from Afro-Caribbean and Caucasian populations. Genomics 1996; 37: 41–50.
Mansur AH, Bishop DT, Holgate ST, Markham AF, Morrison JF . Linkage/association study of a locus modulating total serum IgE on chromosome 14q13-24 in families with asthma. Thorax 2004; 59: 876–882.
Bleecker ER, Amelung PJ, Levitt RC, Postma DS, Meyers DA . Evidence for linkage of total serum IgE and bronchial hyperresponsiveness to chromosome 5q: a major regulatory locus important in asthma. Clin Exp Allergy 1995; 25 (Suppl 2): 84–88; discussion 95–6.
Gobet R, Cerny A, Ruedi E, Hengartner H, Zinkernagel RM . The role of antibodies in natural and acquired resistance of mice to vesicular stomatitis virus. Exp Cell Biol 1988; 56: 175–180.
Parmentier HK, Lammers A, Hoekman JJ, De Vries Reilingh G, Zaanen IT, Savelkoul HF . Different levels of natural antibodies in chickens divergently selected for specific antibody responses. Dev Comp Immunol 2004; 28: 39–49.
Kachamakova NM, Irnazarow I, Parmentier HK, Savelkoul HF, Pilarczyk A, Wiegertjes GF . Genetic differences in natural antibody levels in common carp (Cyprinus carpio L.). Fish Shellfish Immunol 2006; 21: 404–413.
Herzenberg LA, Stall AM, Lalor PA, Sidman C, Moore WA, Parks DR et al. The Ly-1 B cell lineage. Immunol Rev 1986; 93: 81–102.
Lee NJ, Rigby RJ, Gill H, Boyle JJ, Fossati-Jimack L, Morley BJ et al. Multiple loci are linked with anti-red blood cell antibody production in NZB mice—comparison with other phenotypes implies complex modes of action. Clin Exp Immunol 2004; 138: 39–46.
Lu R, Medina KL, Lancki DW, Singh H . IRF-4,8 orchestrate the pre-B-to-B transition in lymphocyte development. Genes Dev 2003; 17: 1703–1708.
Fanzo JC, Hu CM, Jang SY, Pernis AB . Regulation of lymphocyte apoptosis by interferon regulatory factor 4 (IRF-4). J Exp Med 2003; 197: 303–314.
Falini B, Fizzotti M, Pucciarini A, Bigerna B, Marafioti T, Gambacorta M et al. A monoclonal antibody (MUM1p) detects expression of the MUM1/IRF4 protein in a subset of germinal center B cells, plasma cells, and activated T cells. Blood 2000; 95: 2084–2092.
Nishiya N, Yamamoto K, Imaizumi Y, Kohno T, Matsuyama T . Identification of a novel GC-rich binding protein that binds to an indispensable element for constitutive IRF-4 promoter activity in B cells. Mol Immunol 2004; 41: 855–861.
Broman KW, Wu H, Sen S, Churchill GA . R/qtl: QTL mapping in experimental crosses. Bioinformatics 2003; 19: 889–890.
Lander E, Kruglyak L . Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet 1995; 11: 241–247.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001; 25: 402–408.
Acknowledgements
We acknowledge Fundacão para a Ciência e Tecnologia for financial support through Grants no. 62964 and SFRH/BD/29212/2006.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Côrte-Real, J., Rodo, J., Almeida, P. et al. Irf4 is a positional and functional candidate gene for the control of serum IgM levels in the mouse. Genes Immun 10, 93–99 (2009). https://doi.org/10.1038/gene.2008.73
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2008.73