Abstract
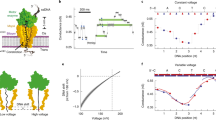
Nanopores can be used to analyse DNA by monitoring ion currents as individual strands are captured and driven through the pore in single file by an applied voltage. Here, we show that serial replication of individual DNA templates can be achieved by DNA polymerases held at the α-haemolysin nanopore orifice. Replication is blocked in the bulk phase, and is initiated only after the DNA is captured by the nanopore. We used this method, in concert with active voltage control, to observe DNA replication catalysed by bacteriophage T7 DNA polymerase (T7DNAP) and by the Klenow fragment of DNA polymerase I (KF). T7DNAP advanced on a DNA template against an 80-mV load applied across the nanopore, and single nucleotide additions were measured on the millisecond timescale for hundreds of individual DNA molecules in series. Replication by KF was not observed when this enzyme was held on top of the nanopore orifice at an applied potential of 80 mV. Sequential nucleotide additions by KF were observed upon applying controlled voltage reversals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Benner, S. et al. Sequence-specific detection of individual DNA polymerase complexes in real time using a nanopore. Nature Nanotech. 2, 718–724 (2007).
Cockroft, S. L., Chu, J., Amorin, M. & Ghadiri, M. R. A single-molecule nanopore device detects DNA polymerase activity with single-nucleotide resolution. J. Am. Chem. Soc. 130, 818–820 (2008).
Hurt, N., Wang, H., Akeson, M. & Lieberman, K. R. Specific nucleotide binding and rebinding to individual DNA polymerase complexes captured on a nanopore. J. Am. Chem. Soc. 131, 3772–3778 (2009).
Wilson, N. A. et al. Electronic control of DNA polymerase binding and unbinding to single DNA molecules. ACS Nano 3, 995–1003 (2009).
Gyarfas, B. et al. Mapping the position of DNA polymerase-bound DNA templates in a nanopore at 5 Å resolution. ACS Nano 3, 1457–1466 (2009).
Song, L. et al. Structure of Staphylococcal alpha-hemolysin, a heptameric transmembrane pore. Science 274, 1859–1866 (1996).
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl Acad. Sci. USA 93, 13770–13773 (1996).
Branton, D. et al. The potential and challenges of nanopore sequencing. Nature Biotechnol. 26, 1146–1153 (2008).
Moffitt, J. R., Chemla, Y. R., Smith, S. B. & Bustamante, C. Recent advances in optical tweezers. Annu. Rev. Biochem. 77, 205–228 (2008).
Lechner, R. L., Engler, M. J. & Richardson, C. C. Characterization of strand displacement synthesis catalyzed by bacteriophage T7 DNA polymerase. J. Biol. Chem. 258, 11174–11184 (1983).
Lechner, R. L. & Richardson, C. C. A preformed, topologically stable replication fork. Characterization of leading strand DNA synthesis catalyzed by T7 DNA polymerase and T7 gene 4 protein. J. Biol. Chem. 258, 11185–11196 (1983).
Tabor, S. & Richardson, C. C. Selective inactivation of the exonuclease activity of bacteriophage T7 DNA polymerase by in vitro mutagenesis. J. Biol. Chem. 264, 6447–6458 (1989).
Canceill, D., Viguera, E. & Ehrlich, S. D. Replication slippage of different DNA polymerases is inversely related to their strand displacement efficiency. J. Biol. Chem. 274, 27481–27490 (1999).
Asseline, U. et al. Nucleic acid-binding molecules with high affinity and base sequence specificity: intercalating agents covalently linked to oligodeoxynucleotides. Proc. Natl Acad. Sci. USA 81, 3297–3301 (1984).
Asseline, U., Bonfils, E., Dupret, D. & Thuong, N. T. Synthesis and binding properties of oligonucleotides covalently linked to an acridine derivative: new study of the influence of the dye attachment site. Bioconjug. Chem. 7, 369–379 (1996).
Patel, S. S., Wong, I. & Johnson, K. A. Pre-steady-state kinetic analysis of processive DNA replication including complete characterization of an exonuclease-deficient mutant. Biochemistry 30, 511–525 (1991).
Huber, H. E., Tabor, S. & Richardson, C. C. Escherichia coli thioredoxin stabilizes complexes of bacteriophage T7 DNA polymerase and primed templates. J. Biol. Chem. 262, 16224–16232 (1987).
Datta, K. & LiCata, V. J. Salt dependence of DNA binding by Thermus aquaticus and Escherichia coli DNA polymerases. J. Biol. Chem. 278, 5694–5701 (2003).
Rothwell, P. J. & Waksman, G. Structure and mechanism of DNA polymerases. Adv. Protein Chem. 71, 401–440 (2005).
Astatke, M., Grindley, N. D. & Joyce, C. M. How E. coli DNA polymerase I (Klenow fragment) distinguishes between deoxy- and dideoxynucleotides. J. Mol. Biol. 278, 147–165 (1998).
Tabor, S. & Richardson, C. C. A single residue in DNA polymerases of the Escherichia coli DNA polymerase I family is critical for distinguishing between deoxy- and dideoxyribonucleotides. Proc. Natl Acad. Sci. USA 92, 6339–6343 (1995).
Akeson, M., Branton, D., Kasianowicz, J. J., Brandin, E. & Deamer, D. W. Microsecond time-scale discrimination among polycytidylic acid, polyadenylic acid and polyuridylic acid as homopolymers or as segments within single RNA molecules. Biophys. J. 77, 3227–3233 (1999).
Meller, A., Nivon, L., Brandin, E., Golovchenko, J. & Branton, D. Rapid nanopore discrimination between single polynucleotide molecules. Proc. Natl Acad. Sci. USA 97, 1079–1084 (2000).
Kibbe, W. A. OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Res. 35, W43–W46 (2007).
Acknowledgements
The authors are grateful to P. Walker at Stanford University PAN for expert oligonucleotide synthesis, Y. Kolodji for help with data analysis, B. Gyarfas for making the Supplementary movies and for assistance with FSM implementation, D. Garalde for advice on the use of T7DNAP, and W. Kibbe (Northwestern University) for modifications to OligoCalc. This work was supported by grants from Oxford Nanopore Technologies and from NHGRI (1RC2HG005553-01) to M.A. and D.D.
Author information
Authors and Affiliations
Contributions
F.O. designed experiments and performed data analysis. K.R.L. co-authored the manuscript, and designed and conducted experiments. S.B., G.M.C. and J.M.D. conducted nanopore experiments, including FSM implementation. D.W.D. conceived the idea of coupling polymerases to nanopores and helped design experiments. M.A. co-authored the manuscript, conceived the blocking oligomer strategy, designed experiments and is responsible for the overall quality of the work.
Corresponding author
Ethics declarations
Competing interests
M. Akeson and D. Deamer are consultants to Oxford Nanopore Technologies (Oxford, England). Oxford Nanopore Technologies has licensed rights to some of the inventions detailed in this manuscript and has sponsored some of the research presented here through a grant to the University of California.
Supplementary information
Supplementary information
Supplementary information (PDF 802 kb)
Supplementary information
Supplementary movie 1 (AVI 5643 kb)
Supplementary information
Supplementary movie 2 (AVI 27245 kb)
Rights and permissions
About this article
Cite this article
Olasagasti, F., Lieberman, K., Benner, S. et al. Replication of individual DNA molecules under electronic control using a protein nanopore. Nature Nanotech 5, 798–806 (2010). https://doi.org/10.1038/nnano.2010.177
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2010.177
This article is cited by
-
Mitogenome-wise codon usage pattern from comparative analysis of the first mitogenome of Blepharipa sp. (Muga uzifly) with other Oestroid flies
Scientific Reports (2022)
-
Single-molecule DNA sequencing using two-dimensional Ti2C(OH)2 MXene nanopores: A first-principles investigation
Nano Research (2022)
-
Revealing the Effects of Pore Size and Geometry on the Mechanical Properties of Graphene Nanopore Using the Atomistic Finite Element Method
Acta Mechanica Solida Sinica (2019)
-
Nanotechnology based approaches for detection and delivery of microRNA in healthcare and crop protection
Journal of Nanobiotechnology (2018)
-
Three decades of nanopore sequencing
Nature Biotechnology (2016)