Abstract
All Gram-negative bacteria, mitochondria and chloroplasts have outer membrane proteins (OMPs) that perform many fundamental biological processes. The OMPs in Gram-negative bacteria are inserted and folded into the outer membrane by the β-barrel assembly machinery (BAM). The mechanism involved is poorly understood, owing to the absence of a structure of the entire BAM complex. Here we report two crystal structures of the Escherichia coli BAM complex in two distinct states: an inward-open state and a lateral-open state. Our structures reveal that the five polypeptide transport-associated domains of BamA form a ring architecture with four associated lipoproteins, BamB–BamE, in the periplasm. Our structural, functional studies and molecular dynamics simulations indicate that these subunits rotate with respect to the integral membrane β-barrel of BamA to induce movement of the β-strands of the barrel and promote insertion of the nascent OMP.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Walther, D. M., Rapaport, D. & Tommassen, J. Biogenesis of β-barrel membrane proteins in bacteria and eukaryotes: evolutionary conservation and divergence. Cell. Mol. Life Sci. 66, 2789–2804 (2009)
Tommassen, J. Assembly of outer-membrane proteins in bacteria and mitochondria. Microbiology 156, 2587–2596 (2010)
Wu, T. et al. Identification of a multicomponent complex required for outer membrane biogenesis in Escherichia coli . Cell 121, 235–245 (2005)
Sasaki, K. et al. VDAC: Old protein with new roles in diabetes. Am. J. Physiol. Cell Physiol. 303, C1055–C1060 (2012)
Bender, A. et al. TOM40 mediates mitochondrial dysfunction induced by α-synuclein accumulation in Parkinson’s disease. PLoS ONE 8, e62277 (2013)
Knowles, T. J., Scott-Tucker, A., Overduin, M. & Henderson, I. R. Membrane protein architects: the role of the BAM complex in outer membrane protein assembly. Nature Rev. Microbiol. 7, 206–214 (2009)
Hagan, C. L., Silhavy, T. J. & Kahne, D. β-barrel membrane protein assembly by the Bam complex. Annu. Rev. Biochem. 80, 189–210 (2011)
Ricci, D. P. & Silhavy, T. J. The Bam machine: a molecular cooper. Biochim. Biophys. Acta 1818, 1067–1084 (2012)
Rigel, N. W. & Silhavy, T. J. Making a β-barrel: assembly of outer membrane proteins in Gram-negative bacteria. Curr. Opin. Microbiol. 15, 189–193 (2012)
Webb, C. T., Heinz, E. & Lithgow, T. Evolution of the β-barrel assembly machinery. Trends Microbiol. 20, 612–620 (2012)
Noinaj, N., Rollauer, S. E. & Buchanan, S. K. The β-barrel membrane protein insertase machinery from Gram-negative bacteria. Curr. Opin. Struct. Biol. 31, 35–42 (2015)
Misra, R., Stikeleather, R. & Gabriele, R. In vivo roles of BamA, BamB and BamD in the biogenesis of BamA, a core protein of the β-barrel assembly machine of Escherichia coli . J. Mol. Biol. 427, 1061–1074 (2015)
McMorran, L. M., Brockwell, D. J. & Radford, S. E. Mechanistic studies of the biogenesis and folding of outer membrane proteins in vitro and in vivo: what have we learned to date? Arch. Biochem. Biophys. 564, 265–280 (2014)
Hagan, C. L., Kim, S. & Kahne, D. Reconstitution of outer membrane protein assembly from purified components. Science 328, 890–892 (2010)
Hagan, C. L., Westwood, D. B. & Kahne, D. Bam lipoproteins assemble BamA in vitro . Biochemistry 52, 6108–6113 (2013)
Roman-Hernandez, G., Peterson, J. H. & Bernstein, H. D. Reconstitution of bacterial autotransporter assembly using purified components. Elife 3, e04234 (2014)
Kim, S. et al. Structure and function of an essential component of the outer membrane protein assembly machine. Science 317, 961–964 (2007)
Noinaj, N. et al. Structural insight into the biogenesis of β-barrel membrane proteins. Nature 501, 385–390 (2013)
Ni, D. et al. Structural and functional analysis of the β-barrel domain of BamA from Escherichia coli . FASEB J. 28, 2677–2685 (2014)
Albrecht, R. et al. Structure of BamA, an essential factor in outer membrane protein biogenesis. Acta Crystallogr. D 70, 1779–1789 (2014)
Noinaj, N., Fairman, J. W. & Buchanan, S. K. The crystal structure of BamB suggests interactions with BamA and its role within the BAM complex. J. Mol. Biol. 407, 248–260 (2011)
Albrecht, R. & Zeth, K. Structural basis of outer membrane protein biogenesis in bacteria. J. Biol. Chem. 286, 27792–27803 (2011)
Dong, C., Yang, X., Hou, H. F., Shen, Y. Q. & Dong, Y. H. Structure of Escherichia coli BamB and its interaction with POTRA of BamA. Acta Crystallogr. D 68, 1134–1139 (2012)
Kim, K. H. & Paetzel, M. Crystal structure of Escherichia coli BamB, a lipoprotein component of the β-barrel assembly machinery complex. J. Mol. Biol. 406, 667–678 (2011)
Heuck, A., Schleiffer, A. & Clausen, T. Augmenting β-augmentation: structural basis of how BamB binds BamA and may support folding of outer membrane proteins. J. Mol. Biol. 406, 659–666 (2011)
Warner, L. R. et al. Structure of the BamC two-domain protein obtained by Rosetta with a limited NMR data set. J. Mol. Biol. 411, 83–95 (2011)
Sandoval, C. M., Baker, S. L., Jansen, K., Metzner, S. I. & Sousa, M. C. Crystal structure of BamD: an essential component of the β-barrel assembly machinery of gram-negative bacteria. J. Mol. Biol. 409, 348–357 (2011)
Dong, C., Hou, H. F., Yang, X., Shen, Y. Q. & Dong, Y. H. Structure of Escherichia coli BamD and its functional implications in outer membrane protein assembly. Acta Crystallogr. D 68, 95–101 (2012)
Knowles, T. J. et al. Structure and function of BamE within the outer membrane and the β-barrel assembly machine. EMBO Rep. 12, 123–128 (2011)
Kim, K. H. et al. Structural characterization of Escherichia coli BamE, a lipoprotein component of the β-barrel assembly machinery complex. Biochemistry 50, 1081–1090 (2011)
Kim, K. H., Aulakh, S. & Paetzel, M. Crystal structure of β-barrel assembly machinery BamCD protein complex. J. Biol. Chem. 286, 39116–39121 (2011)
Jansen, K. B., Baker, S. L. & Sousa, M. C. Crystal structure of BamB bound to a periplasmic domain fragment of BamA, the central component of the β-barrel assembly machine. J. Biol. Chem. 290, 2126–2136 (2015)
Hagan, C. L. & Kahne, D. The reconstituted Escherichia coli Bam complex catalyzes multiple rounds of β-barrel assembly. Biochemistry 50, 7444–7446 (2011)
Rigel, N. W., Ricci, D. P. & Silhavy, T. J. Conformation-specific labeling of BamA and suppressor analysis suggest a cyclic mechanism for β-barrel assembly in Escherichia coli . Proc. Natl Acad. Sci. USA 110, 5151–5156 (2013)
Noinaj, N. & Buchanan, S. K. Structural insights into the transport of small molecules across membranes. Curr. Opin. Struct. Biol. 27, 8–15 (2014)
Noinaj, N., Kuszak, A. J., Balusek, C., Gumbart, J. C. & Buchanan, S. K. Lateral opening and exit pore formation are required for BamA function. Structure 22, 1055–1062 (2014)
Ricci, D. P., Hagan, C. L., Kahne, D. & Silhavy, T. J. Activation of the Escherichia coli β-barrel assembly machine (Bam) is required for essential components to interact properly with substrate. Proc. Natl Acad. Sci. USA 109, 3487–3491 (2012)
Rigel, N. W., Schwalm, J., Ricci, D. P. & Silhavy, T. J. BamE modulates the Escherichia coli β-barrel assembly machine component BamA. J. Bacteriol. 194, 1002–1008 (2012)
Tellez, R., Jr & Misra, R. Substitutions in the BamA β-barrel domain overcome the conditional lethal phenotype of a ΔbamB ΔbamE strain of Escherichia coli. J. Bacteriol . 194, 317–324 (2012)
Bennion, D., Charlson, E. S., Coon, E. & Misra, R. Dissection of β-barrel outer membrane protein assembly pathways through characterizing BamA POTRA 1 mutants of Escherichia coli . Mol. Microbiol. 77, 1153–1171 (2010)
Gessmann, D. et al. Outer membrane β-barrel protein folding is physically controlled by periplasmic lipid head groups and BamA. Proc. Natl Acad. Sci. USA 111, 5878–5883 (2014)
Li, M. Z. & Elledge, S. J. Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nature Methods 4, 251–256 (2007)
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010)
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010)
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012)
Cowtan, K. DM : An automated procedure for phase improvement by density modification. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography 31, 34–38 (1994)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Liu, H. & Naismith, J. H. An efficient one-step site-directed deletion, insertion, single and multiple-site plasmid mutagenesis protocol. BMC Biotechnol. 8, 91 (2008)
Pronk, S. et al. GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29, 845–854 (2013)
de Jong, D. H. et al. Improved parameters for the Martini coarse-grained protein force field. J. Chem. Theory Comput. 9, 687–697 (2013)
Stansfeld, P. J. et al. MemProtMD: Automated insertion of membrane protein structures into explicit lipid membranes. Structure 23, 1350–1361 (2015)
Piggot, T. J., Pineiro, A. & Khalid, S. Molecular dynamics simulations of phosphatidylcholine membranes: a comparative force field study. J. Chem. Theory Comput. 8, 4593–4609 (2012)
Jefferys, E., Sands, Z. A., Shi, J. Y., Sansom, M. S. P. & Fowler, P. W. Alchembed: a computational method for incorporating multiple proteins into complex lipid geometries. J. Chem. Theory Comput. 11, 2743–2754 (2015)
Oostenbrink, C., Villa, A., Mark, A. E. & Van Gunsteren, W. F. A biomolecular force field based on the free enthalpy of hydration and solvation: The GROMOS force-field parameter sets 53A5 and 53A6. J. Comput. Chem. 25, 1656–1676 (2004)
Domański, J., Stansfeld, P. J., Sansom, M. S. P. & Beckstein, O. Lipidbook: a public repository for force-field parameters used in membrane simulations. J. Membr. Biol. 236, 255–258 (2010)
Eswar, N. et al. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinformatics 15, 5.6.1–5.6.30 (2006)
Bussi, G., Zykova-Timan, T. & Parrinello, M. Isothermal-isobaric molecular dynamics using stochastic velocity rescaling. J. Chem. Phys. 130, 074101 (2009)
Parrinello, M. & Rahman, A. Polymorphic transitions in single-crystals - a new molecular-dynamics method. J. Appl. Phys. 52, 7182–7190 (1981)
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008)
Hess, B. GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–437 (2008)
York, D. M., Wlodawer, A., Pedersen, L. G. & Darden, T. A. Atomic-level accuracy in simulations of large protein crystals. Proc. Natl Acad. Sci. USA 91, 8715–8718 (1994)
Michaud-Agrawal, N., Denning, E. J., Woolf, T. B. & Beckstein, O. Software news and updates MDAnalysis: a toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 32, 2319–2327 (2011)
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010)
Suzek, B. E. et al. UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches. Bioinformatics 31, 926–932 (2015)
Biegert, A. & Soding, J. Sequence context-specific profiles for homology searching. Proc. Natl Acad. Sci. USA 106, 3770–3775 (2009)
Acknowledgements
We thank H. D. Bernstein for providing HDB150 strain and plasmid pJH114 and T. J. Silhavy for providing JCM166 cells. We appreciate the staff at I24, I02 and I03 of Diamond Light Source UK for beamtime (proposal mx9475) and their assistance with data collection. C.D. is a recipient of the Wellcome Trust investigator award (WT106121MA), and is supported by Medical research council (G1100110/1). W.W. acknowledges the support of the Science and Technology Program of Guangzhou, China (201510010040) and China National Natural Science Foundation of Guangdong (2015A030313152).
Author information
Authors and Affiliations
Contributions
C.D., W.W. and Y.G. conceived and designed the experiments. C.D. and W.W. supervised the project. Y.G., Y.Ze, Z.W. and Y.Zh designed primers and generated the constructs for protein expression and functional assays. Z.W., Y.G., Y.Ze, H.D., H.L. and Z.Z. participated in expression, purification and crystallization of the BAM complex. Y.G. and Y.Ze expressed the BamACDE complex using the plasmids pJH114, and expressed the BamABCDE complex using the plasmid pYG120. Y.G. and Y.Ze purified and crystallized the complexes, optimized crystallization, obtained well diffracted crystals and performed site-directed mutagenesis, functional assays, heat-modifiability assays and western blot. N.G.P., Y.G., C.D. and W.W. undertook data collection and structure determination of the BamACDE and BamABCDE complexes. P.J.S. performed the molecular dynamics simulations. Y.G., C.D., P.J.S. and N.G.P. prepared tables and figures. C.D., Y.G., W.W., P.J.S. and N.G.P. wrote and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
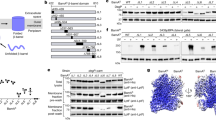
Extended Data Figure 1 BamABCDE and BamACDE complexes and electron density maps contoured at 1σ.
a, Schematic diagram of the five BAM subunits. P-1 to P-5 represent the five BamA POTRA domains. b, SDS–PAGE analysis of the BAM complex from crystals. M, molecular mass marker; 1 and 2 are crystals of the purified BAM complex expressed by construct pYG120 and pJH114, respectively (Supplementary Fig. 1). The BamABCDE crystals contain the full-length BamA–BamE. The crystals were washed five times in fresh reservoir solution, and then dissolved in SDS–PAGE loading buffer. The results showed that the BamB is absent in the BamACDE crystals, while the BamC is complete in both the BamABCDE and BamACDE crystals. c, SDS–PAGE analysis of the purified BAM complex. The BAM complexes expressed from pJH114 is a mixture of BamABCDE and BamACDE complexes (Supplementary Fig. 1). d, 2Fo − Fc electron density map of BamA residues Trp576–Lys580 of BamACDE contoured at 1σ. e, 2Fo − Fc electron density map of BamD residues Tyr177–Trp191 of BamACDE contoured at 1σ. f, 2Fo − Fc electron density map of BamA residues Tyr504–Tyr509 and Phe490–Phe494 of BamABCDE complex contoured at 1σ. g, 2Fo − Fc electron density map of BamB residues Tyr345–Trp348 contoured at 1σ.
Extended Data Figure 2 Superimposition of the two BamACDE complexes in the asymmetric unit.
The BamACDE complex with the full-length BamC, showing BamA (red), BamC (blue), BamD (magenta) and BamE (cyan). Only N-terminal loop of BamC was observed in another BamACDE complex in the asymmetric unit cell (yellow). The structure data suggests that the role of BamC is to retain the ring structure of BamA and BamD during OMP insertion. a, Membrane view of the superimposed BamACDE complexes. The primary difference is one complex has a complete BamC subunit, which binds BamD, BamE, POTRA 1 and 2, while the second complex only the N-terminal coil structure up to Pro88 is observed and the rest of BamC is disordered. The overall structures of the two complexes are very similar with some conformational changes in the β-strands of barrel and extracellular loops with r.m.s.d. of 0.908 Å over 378 Cα atoms, while the periplasmic circular structure has some rotation (see arrows) with a r.m.s.d. of 4.706 Å over 385 Cα atoms. b, Periplasmic view of the superimposition of the two structures. The periplasmic circular structure has some rotations when the C-terminal global domain binds on the POTRA 2. c, Superimposition of the barrels of the two complexes. d, Superimposition of the two BamCs. The N-terminal coil structures superimpose well with a r.m.s.d. of 0.807 Å over 86 Cα atoms.
Extended Data Figure 3 Molecular dynamics simulation of BAM complexes.
a, b, BamABCDE (a) and BamACDE (b) structures modelled with all subunits present and embedded in a model E. coli outer membrane (grey). Phosphate atoms are shown in orange spheres. Lipid-modified cysteine residues of BamB, BamC, BamD and BamE are shown in yellow spheres. c, Both complexes are stable in molecular dynamics simulations, showing limited deviation from the starting configuration (shown in the background). d, Simulations of the complexes of only BamA and BamD subunits retain the ring structure. Without BamC present POTRA 1 (black circle) moves towards the membrane, while POTRA 3 (black arrow) moves towards and interacts with the periplasmic loops of the barrel. The dynamics of POTRA 3 appear to be modulated by BamB. e, Simulations of BamA show enhanced dynamics of the POTRA domains, with POTRA 1 and 2 rotating towards the membrane in an anti-clockwise direction (blue arrow). This separates POTRA 3 from the barrel (black arrow). This conformation of the POTRA domains is unable to form the BAM ring, highlighting the essential nature of BamD and its interactions with BamA in maintaining the ring structure.
Extended Data Figure 4 BamA of the BamABCDE complex is superimposed onto the other published BamA structures.
All the published BamA structures are in the inward-open conformation. In all cases the BamA from BamABCDE is shown in red. a, The BamA of BamABCDE complex is superimposed onto BamA of N. gonorrhoeae (grey) (PDB accession 4K3B)18. The two barrel structures are similar with a r.m.s.d. of 3.803 Å over 385 Cα atoms, but the conformations of the five POTRA domains are quite different. The dotted circle indicates the hydrophobic gap between β1C and β15C. b, BamA of E. coli (magenta) (PDB accession 4N75)19. The two barrel structures superimpose well with a r.m.s.d. of 0.644 Å over 385 Cα atoms, but differences are observed for the β16C terminal residues. The C-terminal residues in BamA of BamABCDE move towards to the lumen of the barrel. c, BamA of E. coli (yellow) (PDB accession 4C4V)20 with a r.m.s.d. of 1.382 Å over 365 barrel Cα atoms. The conformations of the POTRA 5 are quite different. d, BamA of Haemophilus ducreyi (green) (PDB accession 4K3C)18. The barrel structures are similar with a r.m.s.d. of 2.376 Å over 365 barrel Cα atoms, but the conformations of POTRA 4 and 5 are quite different.
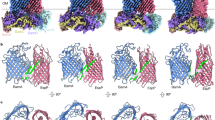
Extended Data Figure 5 The conformational changes between the BamABCDE and BamACDE complexes and heat-modifiability assays of the BamA double cysteine mutants.
The two structures are superimposed onto the BamA barrel structures of BamABCDE and BamACDE complexes with a r.m.s.d. of 4.85 Å over the 379 barrel Cα atoms and a maximum r.m.s.d. of 20 Å. The POTRA domains align with an r.m.s.d. of 5.764 Å over 384 Cα atoms with maximum 15 Å. The BamABCDE complex is in the same colour scheme as Fig. 1. The BamACDE complex is in yellow. The barrel strands β1C–β6C rotate around 65° from BamABCDE to BamACDE, while the BAM periplasmic unit rotates around 30° in a anti-clockwise direction from BamABCDE to BamACDE. a, Membrane view of the superimposition of the BamABCDE and BamACDE complexes. The conformations of BamA POTRA domains, BamB, BamC and BamD are considerably different between the two complexes. b, The periplasmic view of the superimposition of BamABCDE and BamACDE. The circular units rotate around 30° between the two BAM complexes. c, The residues involved in closing the barrel at the periplasmic side in the BamACDE structure. d, Heat-modifiability assays of the BamA double cysteine mutants. SDS–PAGE/western blot analysis of the wild-type BamA, BamA Gly393Cys/Gly584Cys, Glu435Cys/Ser665Cys and Glu435Cys/Ser658Cys mutants showed the heat-modifiability, indicating that the three double cysteine BamA mutants were correctly folded into the OM. F, folded; U, unfolded. See Supplementary Fig. 5.
Extended Data Figure 6 Periplasmic loops bind to BamA POTRA 3, 5, BamD and BamE.
In the BamABCDE complex, the BamA barrel interacts with POTRA 3, 5, BamE and BamD through the periplasmic turns T-1, -2, -3, -5, -6 and -7. a, In the BamABCDE complex, the residues of T-1, -2 and -3 are involved in the interactions with POTRA 5, BamD and BamE. b, Residues in T-5, -6 and -7 interact with POTRA 3 in the BamABCDE complex. c, In the BamACDE complex no interactions are observed between the periplasmic turns and POTRA 3. The figure shows that the residues in T-1, -2 and -3 interact with residues in POTRA 5, BamD and BamE. These structural data may suggest that BamB, C, D and E either directly or indirectly control the conformation of the barrel through its periplasmic turns.
Extended Data Figure 7 BamA and BamD interactions, and superimposition of the BamB structures and the conformational changes of POTRA2 and 3.
a, BamA POTRA 1 and 2 interact with the N-terminal domain of BamD. The interacting residues from both BamA and BamD are shown. b, The functional assays and the western blots were repeated at least three times. Functional assays of the BamA interaction with BamD. The mutation BamA Glu373Lys is lethal, while mutant Arg366Glu impairs the bacterial growth, suggesting these residues may have an important role in the BAM complex. 1–4 are as shown in c. c, Protein expression levels of BamA mutations were detected by western blotting. d, Periplasmic view of BamB of the BamABCDE complex (green) superimposed onto the free BamB structure (orange) (PDB code 3Q7N)21 with a r.m.s.d. of 1.81 Å over 351 Cα atoms with the maximum deviation of 12 Å at loop 19. Loops 15, 19, 23 and 27 of BamB adopt conformational changes to bind to POTRA 2 and 3. e, BamB of the BamABCDE complex superimposed onto BamB in complex with POTRA 3 and 4 (magenta) (PDB code 4PK1)32. The two BamB structures are very similar with a r.m.s.d. of 0.5860 Å over 341 Cα atoms. f, Superimposition of BamABCDE and BamACDE at POTRA 2 and 3 with a r.m.s.d. of 3.57 Å over 159 Cα atoms. In the BamACDE structure the hinge angle between POTRA 2 and 3 is reduced, while POTRA 2 and 3 also separate from BamB, reducing the the interactions between BamB, and POTRA 2 and 3.
Extended Data Figure 8 BamE interacts with BamD and BamC, and BamC interactions with BamD.
BamE interacts with BamA, BamD and BamC. BamC binds extensively to the C-terminal domain of BamD. a, BamE interacts with BamD in the BamACDE complex. BamE contacts the C-terminal domain residues of BamD in the BamACDE complex. b, BamE forms hydrophobic interactions with BamC in the BamACDE complex. BamE residues Pro67, Phe68 and BamC residues Met56 and Ile57 are shown. c, BamC forms contacts with BamA POTRA 1 in BamACDE. BamA residues Phe31, Gln35, Val39 and BamC residues Gly94 and Arg96 are shown. d, BamC interacts with the C-terminal domain of BamD. The interacting residues are shown as sticks. e, BamC interacts with the N-terminal domain of BamD.
Extended Data Figure 9 Conserved residues analysis of BAM complex.
a–d, Consurf residue conservation scores (1–9), plotted onto the molecular surfaces as a colour scale for BamA (red), BamB (green), BamC (blue), BamD (purple) and BamE (cyan), for the BamABCDE structure. Regions of white/grey indicate poorly conserved residues, whereas a more intense colour indicates highly conserved residues. Black dashed circles represent the interaction points on removal of BamC (a), BamD (b), BamE (c) and BamB (d). For each interaction patch a high density of conserved residues is apparent.
Extended Data Figure 10 Conformational differences of the BAM subunits between the BamABCDE and BamACDE complexes, and BAM complex interacts with lipid of the OM.
The subunits of BamABCDE are coloured as in Fig. 1, while the BamACDE subunits are in yellow. a, Superimposition of the BamA subunits onto the barrel domain with an r.m.s.d. of 4.85 Å over the 379 barrel Cα atoms and an r.m.s.d. of 5.76 Å over the 384 Cα atoms of the POTRA domains. The BamA barrel has notable conformational changes in β1C–β6C. The periplasmic POTRA domains rotate about 30° from BamABCDE complex to BamACDE complex, suggesting a novel rotation mechanism to facilitate OMP insertion into the OM. b, Superimposition of the BamC structures. The BamC structures have some conformational changes with a r.m.s.d. of 2.102 Å over 47 Cα atoms of the BamC N-terminal loop. The N-terminal loop Cys25–Val35 becomes more ordered in BamACDE complex. Particularly, the N-terminal domain and the C-terminal domain are ordered and bind to POTRA 1, 2 and the N-terminal domain of BamD in BamACDE complex. The N-terminal loops of the BamC structures superimpose well between residues Val35 and Pro88. c, Superimposition of the BamD structures with an r.m.s.d. of 1.201 Å over 203 Cα atoms. The α-helices are conserved, but the loops have some conformational changes, especially loop 6 (residues Asp121–Asp136) between α-helix 5 and α-helix 6. d, Superimposition of BamE structures with a r.m.s.d. of 1.721 Å over 81 Cα atoms. The β-strands and α-helices of BamE are well conserved, with minor conformational changes observed in the loops. e, Lipid–protein interactions for the BamACDE structure. BamB was modelled into the BamACDE complex by molecular modelling. The BamABCDE complex was built in the OM (Methods), and the residues interacting with lipids of the OM with 4 Å are shown in putty representation to depict lipid interaction residues. Equivalent residues in all five subunits BamA (red), BamB (green), BamC (blue), BamD (purple) and BamE (cyan) interact with the membrane in all three independent simulations. f, Lipid–protein interactions for the BamABCDE structure. BamC was added to the BamABCDE complex by molecular modelling, using the solved domain from the companion complex. BamABCDE complex was inserted into the OMP, with lipid anchors designed (Methods).
Supplementary information
Supplementary Information
This file contains Supplementary Figures and Tables 1-6. (PDF 13747 kb)
Molecular dynamics simulations of BamACDE complex
This video shows the molecular dynamics simulations of BamACDE complex. (MOV 7943 kb)
Side view of BAM machinery from the loading state to the insertion state and the release state
By structurally aligning POTRA 5 from the two X-ray structures it is apparent that periplasmic loops 1 and 2 are very similar in the two structures. Therefore, by taking the first six β-strands from BamACDE and β-strands 7 to 16 from the resting BamABCDE structure we can create a barrel with an open lateral gate between strands β1 and β16. This ‘intermediate-state’ barrel can then be structurally aligned to retain interactions with both POTRA 3 and POTRA 5. This may then be used to illustrate how the rotation and separation of the periplasmic domains may assist the division of the first and last β-strands and thereby promote nascent OMP insertion. (MOV 12312 kb)
Rights and permissions
About this article
Cite this article
Gu, Y., Li, H., Dong, H. et al. Structural basis of outer membrane protein insertion by the BAM complex. Nature 531, 64–69 (2016). https://doi.org/10.1038/nature17199
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature17199
This article is cited by
-
Surveying membrane landscapes: a new look at the bacterial cell surface
Nature Reviews Microbiology (2023)
-
High-throughput screening of BAM inhibitors in native membrane environment
Nature Communications (2023)
-
Peptidoglycan maturation controls outer membrane protein assembly
Nature (2022)
-
HPLC–DAD analysis and antimicrobial activities of Spondias mombin L. (Anacardiaceae)
3 Biotech (2022)
-
HER2-antigen-specific humoral immune response in breast cancer lymphocytes transplanted in hu-PBL hIL-4 NOG mice
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.