Abstract
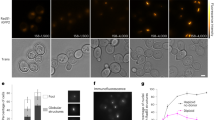
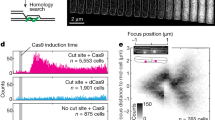
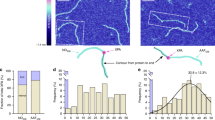
DNA breaks can be repaired with high fidelity by homologous recombination. A ubiquitous protein that is essential for this DNA template-directed repair is RecA1. After resection of broken DNA to produce single-stranded DNA (ssDNA), RecA assembles on this ssDNA into a filament with the unique capacity to search and find DNA sequences in double-stranded DNA (dsDNA) that are homologous to the ssDNA. This homology search is vital to recombinational DNA repair, and results in homologous pairing and exchange of DNA strands. Homologous pairing involves DNA sequence-specific target location by the RecA–ssDNA complex. Despite decades of study, the mechanism of this enigmatic search process remains unknown. RecA is a DNA-dependent ATPase, but ATP hydrolysis is not required for DNA pairing and strand exchange2,3, eliminating active search processes. Using dual optical trapping to manipulate DNA, and single-molecule fluorescence microscopy to image DNA pairing, we demonstrate that both the three-dimensional conformational state of the dsDNA target and the length of the homologous RecA–ssDNA filament have important roles in the homology search. We discovered that as the end-to-end distance of the target dsDNA molecule is increased, constraining the available three-dimensional (3D) conformations of the molecule, the rate of homologous pairing decreases. Conversely, when the length of the ssDNA in the nucleoprotein filament is increased, homology is found faster. We propose a model for the DNA homology search process termed ‘intersegmental contact sampling’, in which the intrinsic multivalent nature of the RecA nucleoprotein filament is used to search DNA sequence space within 3D domains of DNA, exploiting multiple weak contacts to rapidly search for homology. Our findings highlight the importance of the 3D conformational dynamics of DNA, reveal a previously unknown facet of the homology search, and provide insight into the mechanism of DNA target location by this member of a universal family of proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kowalczykowski, S. C., Dixon, D. A., Eggleston, A. K., Lauder, S. D. & Rehrauer, W. M. Biochemistry of homologous recombination in Escherichia coli. Microbiol. Rev. 58, 401–465 (1994)
Menetski, J. P. & Kowalczykowski, S. C. Interaction of recA protein with single-stranded DNA. Quantitative aspects of binding affinity modulation by nucleotide cofactors. J. Mol. Biol. 181, 281–295 (1985)
Kowalczykowski, S. C. & Krupp, R. A. DNA-strand exchange promoted by RecA protein in the absence of ATP: implications for the mechanism of energy transduction in protein-promoted nucleic acid transactions. Proc. Natl Acad. Sci. USA 92, 3478–3482 (1995)
Menetski, J. P., Bear, D. G. & Kowalczykowski, S. C. Stable DNA heteroduplex formation catalyzed by the Escherichia coli RecA protein in the absence of ATP hydrolysis. Proc. Natl Acad. Sci. USA 87, 21–25 (1990)
Adzuma, K. No sliding during homology search by RecA protein. J. Biol. Chem. 273, 31565–31573 (1998)
Kowalczykowski, S. C. Biochemistry of genetic recombination: energetics and mechanism of DNA strand exchange. Annu. Rev. Biophys. Biophys. Chem. 20, 539–575 (1991)
Fulconis, R., Miné, J., Bancaud, A., Dutreix, M. & Viovy, J. L. Mechanism of RecA-mediated homologous recombination revisited by single molecule nanomanipulation. EMBO J. 25, 4293–4304 (2006)
van der Heijden, T. et al. Homologous recombination in real time: DNA strand exchange by RecA. Mol. Cell 30, 530–538 (2008)
Forget, A. L. & Kowalczykowski, S. C. Single-molecule imaging brings Rad51 nucleoprotein filaments into focus. Trends Cell Biol. 20, 269–276 (2010)
McEntee, K., Weinstock, G. M. & Lehman, I. R. Binding of the recA protein of Escherichia coli to single- and double-stranded DNA. J. Biol. Chem. 256, 8835–8844 (1981)
Honigberg, S. M., Gonda, D. K., Flory, J. & Radding, C. M. The pairing activity of stable nucleoprotein filaments made from recA protein, single-stranded DNA, and adenosine 5′-(gamma-thio)triphosphate. J. Biol. Chem. 260, 11845–11851 (1985)
Galletto, R., Amitani, I., Baskin, R. J. & Kowalczykowski, S. C. Direct observation of individual RecA filaments assembling on single DNA molecules. Nature 443, 875–878 (2006)
van Mameren, J. et al. Counting RAD51 proteins disassembling from nucleoprotein filaments under tension. Nature 457, 745–748 (2009)
Julin, D. A., Riddles, P. W. & Lehman, I. R. On the mechanism of pairing of single- and double-stranded DNA molecules by the recA and single-stranded DNA-binding proteins of Escherichia coli. J. Biol. Chem. 261, 1025–1030 (1986)
Gonda, D. K. & Radding, C. M. By searching processively recA protein pairs DNA molecules that share a limited stretch of homology. Cell 34, 647–654 (1983)
Tsang, S. S., Chow, S. A. & Radding, C. M. Networks of DNA and recA protein are intermediates in homologous pairing. Biochemistry 24, 3226–3232 (1985)
Berg, O. G., Winter, R. B. & von Hippel, P. H. Diffusion-driven mechanisms of protein translocation on nucleic acids. 1. Models and theory. Biochemistry 20, 6929–6948 (1981)
Berg, O. G. in The Biology of Nonspecific DNA Protein Interactions (ed. Revzin, A.) 71–85 (CRC, 1990)
Mirshad, J. K. & Kowalczykowski, S. C. Biochemical characterization of a mutant RecA protein altered in DNA-binding loop 1. Biochemistry 42, 5945–5954 (2003)
Harmon, F. G. & Kowalczykowski, S. C. RecQ helicase, in concert with RecA and SSB proteins, initiates and disrupts DNA recombination. Genes Dev. 12, 1134–1144 (1998)
Bianco, P. R. et al. Processive translocation and DNA unwinding by individual RecBCD enzyme molecules. Nature 409, 374–378 (2001)
Perkins, T. T., Quake, S. R., Smith, D. E. & Chu, S. Relaxation of a single DNA molecule observed by optical microscopy. Science 264, 822–826 (1994)
Acknowledgements
We are grateful to members of the laboratory for their comments on this work. A.L.F. was funded by an American Cancer Society Postdoctoral Fellowship (PF-08–046–01-GMC) and S.C.K. was supported by the National Institutes of Health (GM-62653 and GM-64745).
Author information
Authors and Affiliations
Contributions
A.L.F. and S.C.K. conceived the general ideas, designed the experiments and interpreted the data. A.L.F. performed experiments. A.L.F. and S.C.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-4 with legends and legends for Supplementary Movies 1-3. (PDF 531 kb)
Supplementary Movie 1
Composite movie depicting the experimental procedure used to visualize DNA pairing on single DNA-dumbbell molecules by optical trapping (see Supplementary Information file for full legend). (MP4 21540 kb)
Supplementary Movie 2
Movie showing RecA nucleoprotein filaments, both heterologously- and homologously-bound (left and right red spots, respectively) during the extension step (Fig. 2b, step 6) of a pairing assay performed using the 1,762 nt ssDNA (see Supplementary Information file for full legend). (MOV 553 kb)
Supplementary Movie 3
Movie showing RecA nucleoprotein filaments, both heterologously- and homologously-bound (left and right red spots, respectively) during the extension step (Fig. 2b, step 6) of a pairing assay performed using the 430 nt ssDNA (see Supplementary Information file for full legend). (MOV 631 kb)
Rights and permissions
About this article
Cite this article
Forget, A., Kowalczykowski, S. Single-molecule imaging of DNA pairing by RecA reveals a three-dimensional homology search. Nature 482, 423–427 (2012). https://doi.org/10.1038/nature10782
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10782
This article is cited by
-
Cohesin regulates homology search during recombinational DNA repair
Nature Cell Biology (2021)
-
RecA finds homologous DNA by reduced dimensionality search
Nature (2021)
-
Optical tweezers in single-molecule biophysics
Nature Reviews Methods Primers (2021)
-
Transient binding and jumping dynamics of p53 along DNA revealed by sub-millisecond resolved single-molecule fluorescence tracking
Scientific Reports (2020)
-
Promotion of homology-directed DNA repair by polyamines
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.