Host: Benjamin Thompson
Hi, listeners – Benjamin here. It’s a bit of a solo mission for me this week, so we’ve got a slightly shorter show than usual. Don’t worry though, we’ve still got the News Chat – that’s coming up at the end of the show – but first, I’ve got a story about antibiotics and antibiotic resistance.
[Jingle]
Interviewer: Benjamin Thompson
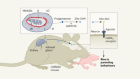
Now, as I’m sure you’re aware, antibiotic resistance is a serious public health issue around the world. As more disease-causing bacteria evolve resistance to antibiotics, researchers are desperately searching for sources of new ones to replace them. And they’re looking in some pretty wild places – inside sea sponges, leafcutter ant colonies and in sediment from the deep ocean bed, to name but three. This week in Nature, a team of scientists have found a new antibiotic and it’s made by bacteria of the genus Photorhabdus, which have a very particular ecological niche, as Kim Lewis from North Eastern University in the US explains.
Interviewee: Kim Lewis
They live in the microbiome or in the gut of nematodes that live in the soil.
Interviewer: Benjamin Thompson
But it gets even more niche than that. These nematodes, which are tiny little worms, are parasites that feed on insect larvae, and they use the Photorhabdus to help. It releases a toxin that paralyses their prey. Both want to eat the larva, but Photorhabdus is not the only bacteria which want to get involved.
Interviewee: Kim Lewis
Of course, there’s now a problem with freeloaders that come from the external environment and also other bacteria that come from the gut of the nematode.
Interviewer: Benjamin Thompson
These freeloading bacteria also want a larval lunch, but Photorhabdus has a trick to fend them off. It releases chemicals that kill off unwanted bacterial competitors. Essentially, it releases antibiotics, and that got Kim’s attention. The chemicals released by Photorhabdus seemed promising as future antibiotics – they’re non-toxic, to the nematodes at least, and they were able to spread through the larval tissue. Strikingly, they also appeared to target Gram-negative bacteria, and these are a type of bacteria that can cause a range of serious diseases from urinary tract infections through to pneumonia, but they can be really hard to treat. Kim and his colleagues found that Photorhabdus was able to kill E. coli – a common Gram-negative bacterium, and so they isolated the compound responsible, which they called darobactin. But there was a problem.
Interviewee: Kim Lewis
It is too big.
Interviewer: Benjamin Thompson
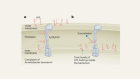
The cell wall of Gram-negative bacteria is made of two membranes, and only small compounds can penetrate them both.
Interviewee: Kim Lewis
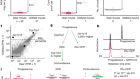
So, we have sort of a rule of thumb for molecules that can penetrate across the outer membrane of Gram-negative bacteria. You need to have a molecular weight of 600 or less, and the molecular weight of the compound we discovered is 960.
Interviewer: Benjamin Thompson
And yet, despite its size, darobactin was still killing E. coli, so how was it doing it? Well, it turns out that darobactin doesn’t need to get inside E. coli at all. Kim found that darobactin’s target is a protein found on the outer membrane of Gram-negatives called BamA, and BamA has been quite a hot topic recently. In fact, two other papers have come out just in the last month describing molecules that target it. And that’s for a reason – BamA plays a really important role.
Interviewee: Kim Lewis
Essentially, it builds the outer membrane, so it takes proteins that the cell synthesises and inserts them into the outer membrane. So, if you jam that protein then you cannot build an outer membrane and the cell dies.
Interviewer: Benjamin Thompson
Kim showed that darobactin was effective in mice, clearing them of infections caused by serious human pathogens, including strains resistant to other antibiotics. Now, Kim’s findings are still quite preliminary, but they caught the attention of Timothy Walsh from Cardiff University, who wasn’t involved in this research.
Interviewee: Timothy Walsh
It’s always good to have a novel compound and this is a novel compound – something we haven’t seen before – so that’s a big tick in the box. The in vivo experimentation seems to be not only promising but very exciting, but it’s early days. There are lots of additional experiments that have to be done before we can start putting this into phase 1 clinical trials.
Interviewer: Benjamin Thompson
Timothy says there are some things to follow up on, such as more extensive toxicity testing and testing darobactin against a wider range of disease-causing bacteria. There have been other curious findings to investigate as well. Kim did isolate E. coli that were resistant to darobactin, but the resistance seems to come at a cost, and although they didn’t die in the presence of darobactin, these bacteria lost their ability to cause disease, for reasons that are currently unclear. Kim is hoping to test more types of darobactin, which he and his team have found encoded in the genomes of other bacteria. Time will tell if darobactin will become part of our clinical armoury, but it’s certainly one to watch, and you can read his paper over at nature.com.
Host: Benjamin Thompson
Coming up, we’ve got an extended News Chat, where I’ll be hearing about the role that chance can play when awarding grants. Next up on the show though, it’s time for the Research Highlights, read this week by Anna Nagle.
[Jingle]
Anna Nagle
Health tracking has become pretty commonplace. You might have a pedometer on your smartphone or a watch that measures your heartrate, but what about a toilet that monitors your metabolism? A team of researchers have been working towards just that, but first they need to find out what information can be extracted from urine. To test this, two of the researchers collected their own urine over ten days. By analysing their fluids, they were able to identify metabolic signatures. In this pilot test, the scientists were able to precisely measure markers of exercise and sleep, along with their coffee and alcohol intake. The researchers believe this first test case shows the potential of urine as a health metric, and eventually hope to create a ‘smart-toilet’ that monitors it. Study that research on your next toilet break over in Nature Digital Medicine.
[Jingle]
Anna Nagle
The smell of old books may be pleasant to some, but for preservationists it could be the tell-tale sign that a book is on its way to the great library in the sky. But now, researchers have developed an electronic nose to sniff out books prone to decay. By collecting a range of books published from 1567 to 2016, the team of scientists were able to identify the compounds emitted by different types of tome. The artificial nose developed by the researchers had six different sensors that picked up these different compounds, allowing it to distinguish between the distinct materials the books were made from and their age. These tell-tale signs suggest whether a book is more or less prone to degradation, so the researchers hope that this sensitive sniffer could help find books in need of preservation. Bookmark that research over in ACS Sensors.
[Jingle]
Interviewer: Benjamin Thompson
Well, finally on this show then, it’s time for the News Chat, and well, it’s an extended News Chat this week, and I am joined by not one but two guests, and I have Nisha Gaind, our European Bureau Chief, and I have Ewen Callaway, a senior reporter here at Nature. Thank you both so much for joining me today. For our first story, we’re going to look at the world of funding, and I’m sure as many of our listeners know, if you apply for a grant, you have to fill in an absolute ream of paperwork and send it off and then you get graded and the top 1% might get funded and maybe the rest of them don’t necessarily make the cut. There’s been a new method of awarding grants put forward which is perhaps a little bit different and a little bit odd. Nisha, what can you tell me about this one?
Interviewee: Nisha Gaind
Yeah, that’s right. This is kind of a cool story and it’s a pretty radical suggestion in the world of funding, as you said. It’s the idea of assigning grants through, effectively, a lottery.
Interviewer: Benjamin Thompson
But how on Earth would you go about divvying up pots of scientific money this way?
Interviewee: Nisha Gaind
In the usual process, as you mention, there is a lot of time spent writing applications and grant review panels ranking these applications and then deciding which ones to fund on the basis of merit. In these new systems that we’re seeing, there is a little bit of initial scanning that is done to ensure that applications meet some kind of quality threshold, and then in some cases, these applications are pretty much picked out of a hat. In fact, one example of a funder says they actually have a hat. In other cases, it’s random number generation. But yeah, it’s as a lottery works.
Interviewer: Benjamin Thompson
So, this is actually being done at the moment. This isn’t some hypothetical thing?
Interviewee: Nisha Gaind
Yeah, that’s right. We’ve got a couple of examples of agencies that are doing this. There’s one in Switzerland – The Swiss National Science Foundation has tried this system with one of its grant types, as have a few funders in New Zealand, and a private foundation, the Volkswagen Foundation, which is Germany’s biggest science funder, has also been using this.
Interviewer: Benjamin Thompson
Presumably then there are some benefits to this system?
Interviewee: Nisha Gaind
Proponents of this method say it just cuts the amount of time that is spent looking at grant applications, and the other is that the random nature of it roots out bias. There are lots of studies that suggest that gender bias and racial bias plagues grant review and peer review and this is one way of rooting out that bias.
Interviewer: Benjamin Thompson
Which seem like good things, certainly, but if I’d spent a month filling in my form and then losing out on the roll of a dice or whatever, I might be a bit miffed. Is everybody happy with this potential way of doing things?
Interviewee: Nisha Gaind
Well, no, and interestingly, we spoke to one person who was actually a recipient of one of these lottery grants and he said that he prefers the original method. He says that researchers put time into writing high-quality applications and that has a lot of value. So, it’s something that we’re seeing at a couple of funders and there is a group of proponents of this method meeting this week to try and encourage others in the research community to try it, but at the moment, it’s still, of course, a minority of funders that are using it.
Interviewer: Benjamin Thompson
Well, let’s move on to our second story today then and Ewen, it’s one of yours, and it is an enormous project looking at the genomes of butterfly species in the US and Canada.
Interviewee: Ewen Callaway
Yeah, so the story I’ve written is about a lab in Texas and they’re collaborators that have decided to sequence the genome of every single butterfly species in the US and Canada: 845 of them, to be precise.
Interviewer: Benjamin Thompson
Which, on the face of it, seems like an enormous undertaking. I mean why do they want to do this?
Interviewee: Ewen Callaway
Yeah, it is an enormous undertaking. I mean DNA sequencing costs have plummeted in recent years, so it’s not that outlandish for a well-funded group because this one is. The collection, I think, is the trickier issue, working with museums and amateur collectors and I think they even took some road trips. Instead of flying to a conference, they just collected butterflies along the way. The reason for doing this, I think, is to study evolution at large. Maybe the traditional approach is to study maybe one or two organisms or a group of organisms, say one genus of butterfly, for instance, and look at patterns of evolution there, but this group decided to look at lots of different groups and see if you can find repeating trends in evolution.
Interviewer: Benjamin Thompson
And, well, what have they found thus far then?
Interviewee: Ewen Callaway
They’re interested in a question, I think, Darwin probably asks, why are there so many animals of one kind and not so many of others? What explains the patterns of biodiversity that we see in the world? And one of their initial insights is, it’s kind of an emerging theme in evolutionary biology, that there’s this kind of link between interbreeding between distantly related species or between distinct species and speciation. So, they’re finding that the groups of species that are the most diverse are also the most likely to have interbred in their past, and that’s a theme we’re seeing with other butterflies, with fish, with Darwin’s finches and other organisms. We’re seeing this tight link between interbreeding and speciation. It’s kind of exciting to see these patterns occurring over and over in the tree of life.
Interviewer: Benjamin Thompson
I mean you say they’ve looked at 845 species, so all of them in the US and in Canada, but presumably there are thousands or hundreds of thousands of species of butterfly across the world. Where does this work sort of fit in and what are they planning to do next?
Interviewee: Ewen Callaway
I don’t know if there are hundreds of thousands of butterflies – there are lots. Yesterday, actually, I was at the Natural History Museum’s insect collection and I went into a room where an expert on butterflies and moths who actually works with this team, he was sorting through this Victorian era collection, this really old collection, and I asked him tongue in cheek: ‘So, how many new species have you found today?’ And he looks around and he’s just like, ‘Well, three, but then I merged two species, so my net total for the day is two.’ But I think that brings home a point that there is just an immense diversity of insects, especially butterflies and moths, and 845 might seem like a lot but it is a drop in the bucket. And there’s this ongoing project called the Earth BioGenome Project which is made up of lots of smaller projects to sequence, ideally, every eukaryote, which are animals, plants, fungi and protozoa, on Earth, and they’ve targeted 1.5 million that we know, but what about all the ones we don’t know? And that’s going to be lots of them, so people have a lot of work to do, and it’s mostly in the collections department.
Interviewer: Benjamin Thompson
Well, finally on this one then Ewen, you mentioned a collections department, you mentioned you going to the Natural History Museum, I mean, did a lot of these genome species then come from sort of going out with your sort of Victorian butterfly net and catching stuff or were they from samples that had been stored away since antiquity or whatever?
Interviewee: Ewen Callaway
It was both. This research group has forged really strong ties with insect collections, probably all over the world, but I think they worked in particular with museums and collections in America, and they’ve got it such that for especially the lower quality genomes that they’re generating for the vast majority of these species, that they can just take one leg and get enough data for a draft genome sequence. Insects have, what, six legs, so if you want a higher quality genome, you’re probably going to need to get some butterfly nets and get an animal and preserve it. It’s a huge undertaking to do the collection but I think this group, like others, has leaned on the collections that have happened over centuries and natural history museums are a real trove for this kind of work.
Interviewer: Benjamin Thompson
Well, Ewen and Nisha, let’s move on to our final story then in this extended News Chat, and let’s stick with DNA, but this time we’re going to be talking about environmental DNA. Now, which one of you would like to explain to me what that is?
Interviewee: Ewen Callaway
I mean, environmental DNA, as the name suggests, is DNA in the environment. It’s this idea that every organism is sluffing off bits of DNA in its cells or what have you and just leaving them in places, so there’s probably a lot of environmental DNA in this recording studio. And the innovation in the last, say, decade and a half has been that this DNA sticks around for a long time and we can just collect it, whether in soil or pond water, whatever, and get an idea of what animals or plants or whatever has been there in the past – that’s basically what environmental DNA is.
Interviewer: Benjamin Thompson
So, it’s kind of a genetic silhouette then of what’s existed in a place at a particular time, and what sort of things has it been used for?
Interviewee: Nisha Gaind
So, in this case, we have an example of a really cool study that has tracked some remote waterholes in Australia, and they’ve been using this technique of tracking environmental DNA to detect an endangered bird for the first time. It’s a bird called a Gouldian finch, and as Ewen said, it’s highlighting the power of this technique using environmental DNA to identify or detect these kind of difficult to track species, and it’s being increasingly used by environmental agencies.
Interviewer: Benjamin Thompson
I guess the advantage is then that if it’s a rare bird, you don’t have to go out with your binoculars and see it – you can see evidence that it was there. But DNA, as we know, is kind of fragile, right, and it can be difficult to detect. There must be some downsides to this particular way of working.
Interviewee: Ewen Callaway
Yeah, I mean I think as Nisha notes, there aren’t a ton of agreed upon standards for doing this sort of work, and I think that’s something that agencies are working on because they see the huge potential of it. One other, I guess you could call it downside, is that you can definitely determine whether an animal is there or not, and maybe by measuring diversity of DNA you can get some idea of that there are multiple animals but unlike traditional surveys of biodiversity, you don’t really get a great census on population numbers, and it’s not an unachievable ask but I think it’s going to be one of the major challenges facing this technology is how do we know how many of something is there, which is really important for all sorts of questions that people might want to ask. But finding rare organisms, I think it’s great. One of my favourite papers came out a couple of years ago and they used environmental DNA in the ocean to census whale sharks, which are the biggest fish in the ocean but they’re just really hard to spot, they’re really rare to see and they congregate in these large groups but we don’t always see them, so it’s quite an exciting application, I think.
Interviewer: Benjamin Thompson
Well, it seems then like you do need a sort of healthy mix of conventional methods and maybe this sort of DNA-based stuff, but what might be the future of eDNA use?
Interviewee: Nisha Gaind
So, we’re seeing this technique increasingly being used and alongside conventional methods but it’s something that scientists say that for both animals and researchers can be more cost-effective and, in some cases, safer than the conventional methods of tracking animals.
Interviewer: Benjamin Thompson
Well, thank you both for joining me today, Nisha and Ewen. Listeners, head over to nature.com/news for more on these stories and more from the wide world of science. And that’s it for this week’s show. We’ll be back next week with another edition of the Nature Podcast. So, until then, I’ve been Benjamin Thompson. Thanks for listening.

 Science funders gamble on grant lotteries
Science funders gamble on grant lotteries
 Every butterfly in the United States and Canada now has a genome sequence
Every butterfly in the United States and Canada now has a genome sequence
 Rare bird’s detection highlights promise of ‘environmental DNA’
Rare bird’s detection highlights promise of ‘environmental DNA’