Abstract
Aim:
To characterize enzymatic activity of severe acute respiratory syndrome (SARS) coronavirus (CoV) 3C-like protease (3CLpro) and its four site-directed mutants.
Methods:
Based on the fluorescence resonance energy transfer (FRET) principle using 5-[(2′-aminoethyl)-amino] naphthelenesulfonic acid (EDANS) and 4-[[4-(dimethylamino) phenyl] azo] benzoic acid (Dabcyl) as the energy transfer pair, one fluorogenic substrate was designed for the evaluation of SARS-CoV 3CLpro proteolytic activity.
Results:
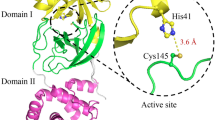
The kinetic parameters of the fluorogenic substrate have been determined as Km=404 μmol·L−1, kcat=1.08 min−1, and kcat/Km=2.7 mmol−1·L·min−1. SARS-CoV 3CLpro showed substantial pH and temperature-triggered activity switches, and site-directed mutagenesis analysis of SARS-CoV 3CLpro revealed that substitutions of His41, Cys145, and His163 resulted in complete loss of enzymatic activity, while replacement of Met162 with Ala caused strongly increased activity.
Conclusion:
This present work has provided valuable information for understanding the catalytic mechanism of SARS-CoV 3CLpro. This FRET-based assay might supply an ideal approach for the exploration SARS-CoV 3CLpro putative inhibitors.
Similar content being viewed by others
Article PDF
References
Holmes KV . SARS coronavirus: a new challenge for prevention and therapy. J Clin Invest 2003; 111: 1605–9.
Peiris JS, Lai ST, Poon LL, Guan Y, Yam LY, Lim W, et al. Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 2003; 361: 1319–25.
Fouchier RA, Kuiken T, Schutten M, Van Amerongen G, Van Doornum GJ, Van Den Hoogen BG, et al. Aetiology: Koch's postulates fulfilled for SARS virus. Nature 2003; 423: 240–7.
Holmes KV . SARS-associated coronavirus. N Engl J Med 2003; 348: 1948–51.
Ziebuhr J, Heusipp G, Siddell SG . Biosynthesis, purification, and characterization of the human coronavirus 229E 3C-like proteinase. J Virol 1997; 71: 3992–7.
Dougherty WG, Semler BL . Expression of virus-encoded proteinases: functional and structural similarities with cellular enzymes. Microbiol Rev 1993; 57: 781–822.
Eleouet JF, Rasschaert D, Lambert P, Levy L, Vende P, Laude H . Complete sequence (20 kilobases) of the polyprotein-encoding gene 1of transmissible gastroenteritis virus. Virology 1995; 206: 817–22.
Thiel V, Herold J, Schelle B, Siddell SG . Viral replicase gene products suffice for coronavirus discontinuous transcription. J Virol 2001; 75: 6676–81.
Herold J, Raabe T, Schelle-Prinz B, Siddell SG . Nucleotide sequence of the human coronavirus 229E RNA polymerase locus. Virology 1993; 195: 680–91.
Lee HJ, Shieh CK, Gorbalenya AE, Koonin EV, Monica N, Tuler J, et al. The complete sequence (22 kilobases) of murine coronavirus gene 1encoding the putative proteases and RNA polymerase. Virology 1991; 180: 567–82.
Liu DX, Brown TD . Characterization and mutational analysis of an ORF 1a-encoding proteinase domain responsible for proteolytic processing of the infectious bronchitis virus 1a/1b polyprotein. Virology 1995; 209: 420–7.
Ziebuhr J, Snijder EJ, Gorbalenya AE . Virus-encoded proteinases and proteolytic processing in Nidovirales. J Gen Virol 2000; 81: 853–79.
Hegyi A, Ziebuhr J . Conservation of substrate specificities among coronavirus main proteases. J Gen Virol 2002; 83: 595–9.
Ziebuhr J, Herold J, Siddell SG . Characterization of a human coronavirus (strain 229E) 3C-like proteinase activity. J Virol 1995; 69: 4331–8.
Anand K, Plam GJ, Mesters JR, Siddell SG, Ziebuhr J, Higenfeld R . Structure of coronavirus main proteinase reveals combination of a chymotrypsin fold with an extra alpha-helical domain. EMBO J 2002; 21: 3213–24.
Fan KQ, Wei P, Feng Q, Chen SD, Huang CK, Ma L, et al. Biosynthesis, purification, and substrate specificity of severe acute respiratory syndrome coronavirus 3C-like proteinase. J Biol Chem 2004; 279: 1637–42.
Anand K, Ziebuhr J, Wadhwani P, Mesters JR, Hilgenfeld R . Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs. Science 2003; 300: 1763–7.
Yang HT, Yang MJ, Ding Y, Liu YW, Lou ZY, Zhou Z, et al. The crystal structures of severe acute respiratory syndrome virus main protease and its complex with an inhibitor. Proc Natl Acad Sci USA 2003; 100: 13190–5.
Kim JC, Spence RA, Currier PF, Liu X, Denison MR . Coronavirus protein processing and RNA synthesis is inhibited by the cysteine protease inhibitor E43d. Virology 1995; 208: 1–8.
Someya Y, Takeda N, Miyamura T . Identification of active-site amino acid residues in the Chiba virus 3C-like protease. J Virol 2002; 76: 5949–58.
Xiong B, Gui CS, Xu XY, Luo C, Chen J, Luo HB, et al. A 3D model of SARS-CoV 3CL proteinase and its inhibitors design by virtual screening. Acta Pharmacol Sin 2003; 24: 497–504.
Sun HF, Luo HB, Yu CY, Sun T, Chen J, Peng SY, et al. Molecular cloning, expression, purification, and mass spectrometric characterization of 3C-like protease of SARS coronavirus. Protein Exp Purif 2003; 32: 302–8.
Knight CG, Willenbrock F, Murphy G . A novel coumarin-labelled peptide for sensitive continuous assays of the matrix metalloproteinases. FEBS Lett 1992; 296: 263–6.
Angliker H, Neumann U, Molloy SS, Thomas G . Internally quenched fluorogenic substrate for furin. Anal Biochem 1995; 224: 409–12.
Mittoo S, Sundstrom LE, Bradley M . Synthesis and evaluation of fluorescent probes for the detection of calpain activity. Anal Biochem 2003; 319: 234–8.
Kuo CJ, Chi YH, Hsu JT, Liang PH . Characterization of SARS main protease and inhibitor assay using a fluorogenic substrate. Biochem Biophys Res Commun 2004; 318: 862–7.
Garcia-Echeverria C, Rich DH . New intramolecularly quenched fluorogenic peptide substrates for the study of the kinetic specificity of papain. FEBS Lett 1992; 297: 100–2.
Wang GT, Matayoshi E, Jan Huffaker H, Krafft GA . Design and synthesis of new fluorogenic HIV protease substrates based on resonance energy transfer. Tetrahedron Lett 1990; 31: 6493–6.
Matayoshi ED, Wang GT, Krafft GA, Erickson J . Novel fluorogenic substrates for assaying retroviral proteases by resonance energy transfer. Science 1990; 247: 954–8.
Maggiora LL, Smith CW, Zhang ZY . A general method for the preparation of internally quenched fluorogenic protease substrates using solid-phase peptide synthesis. J Med Chem 1992; 35: 3727–30.
Huang CK, Wei P, Fan KQ, Liu Y, Lai LL . 3C-like proteinase from SARS coronavirus catalyzes substrate hydrolysis by a general base mechanism. Biochemistry 2004; 43: 4568–74.
Copeland RA . Enzymes: A Practical Introduction to Structure, Mechanism, and Data Analysis. 2nd ed. New York: Wiley-VCH Inc; 2000.
Gorbalenya AE, Koonin EV, Donchenko AP, Blinov VM . Coronavirus genome: prediction of putative functional domains in the non-structural polyprotein by comparative amino acid sequence analysis. Nucleic Acids Res 1989; 17: 4847–61.
Marra MA, Jones SJM, Astell CR, Holt RA, Wilson AB, Butterfield YSN, et al. The genome sequence of the SARS-associated coronavirus. Science 2003; 300: 1399–403.
Lu Y, Denison MR . Determinants of mouse hepatitis virus 3C-like proteinase activity. Virology 1997; 230: 335–42.
Hegyi A, Friebe A, Gorbalenya AE, Ziebuhr J . Mutational analysis of the active centre of coronavirus 3C-like proteases. J Gen Virol 2002; 83: 581–93.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Project supported by the State Key Program of Basic Research of China (grants 2003-CB514125, 2003CB514124, 2002CB512807, 2002CB512802, 2002AA233011), Sino-European Project on SARS Diagnostics and Antivirals (Proposal/Contract No 003831), and the special programs of oppugning SARS from the Ministry of Science and Technology, Chinese Academy of Sciences, National Natural Science Foundation of China and Shanghai Science and Technology Commission.
Rights and permissions
About this article
Cite this article
Chen, S., Chen, Ll., Luo, Hb. et al. Enzymatic activity characterization of SARS coronavirus 3C-like protease by fluorescence resonance energy transfer technique . Acta Pharmacol Sin 26, 99–106 (2005). https://doi.org/10.1111/j.1745-7254.2005.00010.x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1111/j.1745-7254.2005.00010.x
Keywords
This article is cited by
-
Spectroscopic methods for COVID-19 detection and early diagnosis
Virology Journal (2022)
-
Artificial intelligence method for predicting the maximum stress of an off-center casing under non-uniform ground stress with support vector machine
Science China Technological Sciences (2020)
-
Kinetic characterization of trans-proteolytic activity of Chikungunya virus capsid protease and development of a FRET-based HTS assay
Scientific Reports (2015)
-
Discovery of novel aromatase inhibitors using a homogeneous time-resolved fluorescence assay
Acta Pharmacologica Sinica (2014)