Abstract
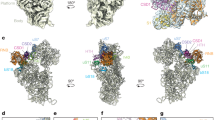
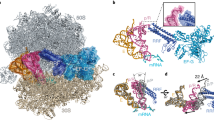
UNDERSTANDING the process whereby the ribosome translates the genetic code into protein molecules will ultimately require high-resolution structural information, and we report here the first crystal structure of a protein from the small ribosomal subunit. This protein, S5, has a molecular mass of 17,500 and is highly conserved in all lifeforms1–4. The molecule contains two distinct α/β domains that have structural similarities to several other proteins that are components of ribonucleoprotein complexes. Mutations in S5 result in several phenotypes which suggest that S5 may have a role in translational fidelity and translocation. These include ribosome ambiguity or ram5, reversion from streptomycin dependence6 and resistance to spectinomycin6. Also, a cold-sensitive, spectinomycin-resistant mutant of S5 has been identified which is defective in initiation7. Here we show that these mutations map to two distinct regions of the molecule which seem to be sites of interaction with ribosomal RNA. A structure/function analysis of the molecule reveals discrepancies with current models8,9 of the 308 subunit.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Wittmann-Liebold, B. & Greuer, B. FEBS Lett. 95, 91–98 (1978).
Kimura, M. J. biol. Chem. 259, 1051–1055 (1984).
Scholzen, T. & Arndt, E. Molec. gen. Genet. 228, 70–80 (1991).
All-Robyn, J. A., Brown, N., Otaka, E. & Liebman, S. W. Molec. cell. Biol. 10, 6544–6553 (1990).
Piepersberg, W., Böck, A. & Wittmann, H.-G. Molec. gen. Genet. 140, 91–100 (1975).
Piepersberg, W., Böck, A., Yaguchi, M. & Wittmann, H.-G. Molec. gen. Genet. 143, 43–52 (1975).
Nomura, M. Cold Spring Harb. Symp. quant. Biol. 52, 653–663 (1987).
Stern, S., Weiser, B. & Noller, H. F. J. molec. Biol. 204, 447–481 (1988).
Brimacombe, R., Atmadja, J., Stiege, W. & Schüler, D. J. molec. Biol. 199, 115–136 (1988).
Appelt, K., White, S. W. & Wilson, K. S. J. biol. Chem. 258, 13328–13330 (1983).
Ramakrishnan, V. & Gerchman, S. E. J. biol. Chem. 266, 880–885 (1991).
White, S. W., Appelt, K., Dijk, J. & Wilson, K. S. FEBS Lett. 163, 73–75 (1983).
Leijonmarck, M. & Liljas, A. J. molec. Biol. 195, 555–580 (1987).
Wilson, K. S., Appelt, K., Badger, J., Tanaka, I. & White, S. W. Proc. natn. Acad. Sci. U.S.A. 83, 7251–7255 (1986).
Hoffmann, D. W., Query, C. C., Golden, B. L., White, S. W. & Keene, J. D. Proc. natn. Acad. Sci. U.S.A. 88, 2495–2499 (1991).
Nagai, K., Oubridge, C., Jessen, T. H., Li, J. & Evans, P. R. Nature 348, 515–520 (1990).
Leijonmarck, M. et al. Proteins 3, 243–251 (1988).
Moazed, D. & Noller, H. F. Nature 327, 389–394 (1987).
Brimacombe, R. Biochimie 73, 927–936 (1991).
Montandon, P. E., Wagner, R. & Stutz, E. EMBO J. 5, 3705–3708 (1986).
Sigmund, C., Ettayebi, M. & Morgan, E. A. Nucleic Acids Res. 12, 4653–4663 (1984).
Ollis, D. L. & White, S. W. Chem. Rev. 87, 981–995 (1987).
Noller, H. F. A. Rev. Biochem. 60, 191–227 (1991).
Capel, M. S. et al. Science 238, 1403–1406 (1987).
Lambert, J. M., Boileau, G., Cover, J. A. & Traut, R. R. Biochemistry 22, 3913–3920 (1983).
Osswald, M. et al. Nucleic Acids Res. 15, 3221–3240 (1987).
Stern, S., Powers, T., Changchien, L.-M. & Noller, H. F. Science 244, 783–790 (1989).
Held, W. A., Mizushima, S. & Nomura, M. J. biol. Chem. 248, 5720–5730 (1973).
Dontsova, O., Kopylov, A. & Brimacombe, R. EMBO J. 10, 2613–2620 (1991).
Arndt, U. W. Meth. Enzym. 114, 472–485 (1985).
Furey, W. & Swaminathan, S. Am. Crystallogr. Assn. Abstr. Ser. 2, Vol. 18, 73 (1990).
Jones, T. A., Zou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Acta Crystallogr. A47, 110–119 (1991).
Brünger, A. T. J. molec. Biol. 203, 803–816 (1988) (Program X-PLOR, Yale Univ., 1989).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ramakrishnan, V., White, S. The structure of ribosomal protein S5 reveals sites of interaction with 16S rRNA. Nature 358, 768–771 (1992). https://doi.org/10.1038/358768a0
Issue Date:
DOI: https://doi.org/10.1038/358768a0
This article is cited by
-
Spontaneous spectinomycin resistance mutations detected after biolistic transformation of Daucus carota L.
Physiology and Molecular Biology of Plants (2011)
-
Spectinomycin Resistance in rpsE Mutants is Recessive in Streptomyces roseosporus
The Journal of Antibiotics (2005)
-
Structure of the 30S ribosomal subunit
Nature (2000)
-
Structure of a bacterial 30S ribosomal subunit at 5.5 Å resolution
Nature (1999)
-
Site-specific DNA binding using a variation of the double stranded RNA binding motif
Nature Structural Biology (1998)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.