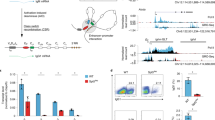
Abstract
Transcription of the immunoglobulin κ light-chain genes depends on the presence of a TATA box upstream of the leader gene segment1,2 and is regulated by an enhancer sequence in the large intron3,4. In studying a rearranged mouse κ light-chain gene we have now found that sequences between −90 and −160 base pairs (bp) upstream of the coding region are essential for correct transcription in gene transfer experiments. This region contains the deca- and pentadecanucleotide sequences TNATTTGCAT and TGCAGCTGTGNCCAG, which we call dc and pd, respectively. Sequences related to dc and pd were found upstream of all human and mouse κ-chain variable region (Vκ) genes, upstream of λ-chain variable region (Vλ) genes, and within the mouse heavy-chain enhancer5,6. An inverted and complementary form of the dc element (ATGCAAATNA, called cd) occurs upstream of all heavy-chain variable region (VH) genes. The newly defined sequences may be involved in the control of immunoglobulin gene transcription.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Kelley, D. E., Coleclough, C. & Perry, R. P. Cell 29, 681–689 (1982).
Bentley, D. L., Farrell, P. J. & Rabbitts, T. H. Nucleic Acids Res. 10, 1841–1856 (1982).
Picard, D. & Schaffner, W. Nature 307, 80–82 (1984).
Queen, C. & Baltimore, D. Cell 33, 741–748 (1983).
Banerji, J., Olson, L. & Schaffner, W. Cell 33, 729–740 (1983).
Gillies, S. D., Morrison, S. L., Oi, V. & Tonegawa, S. Cell 33, 717–728 (1983).
Falkner, F. G. & Zachau, H. G. Nature 298, 286–288 (1982).
Gillies, S. D. & Tonegawa, S. Nucleic Acids Res. 11, 7981–7997 (1983).
Stafford, J. & Queen, C. Nature 306, 77–79 (1983).
Mulligan, R. C. & Berg, P. Proc. natn. Acad. Sci. U.S.A. 78, 2072–2076 (1981).
Altenburger, W., Steinmetz, M. & Zachau, H. G. Nature 287, 603–607 (1980).
Kearney, J. F., Liesegang, B. & Rajewsky, K. J. Immun. 123, 1548–1550 (1979).
Neumann, E., Schaefer-Ridder, M., Wang, Y. & Hofschneider, P. H. EMBO J. 1, 841–845 (1982).
Berk, A. J. & Sharp, P. A. Cell 12, 721–732 (1977).
Köhler, G., Howe, S. C. & Milstein, C. Eur. J. Immun. 6, 292–295 (1976).
Heinrich, G., Traunecker, A. & Tonegawa, S. J. exp. Med. 159, 417–435 (1984).
Benoist, C. & Chambon, P. Nature 290, 304–310 (1981).
Gruss, P., Dhar, R. & Khoury, G. Proc. natn. Acad. Sci. U.S.A. 78, 943–947 (1981).
Igo-Kemenes, T., Hörz, W. & Zachau, H. G. A. Rev. Biochem. 51, 89–121 (1982).
Weischet, W. O., Glotov, B., Schnell, H. & Zachau, H.G. Nucleic Acids Res. 10, 3627–3645 (1982).
Mills, F. C., Fisher, M. L., Kuroda, R., Ford, A. M. & Gould, J. H. Nature 306, 809–812 (1983).
Falkner, F. G. thesis, Univ. München (1984).
Davidson, E. H., Jacobs, H. T. & Britten, R. J. Nature 301, 468–470 (1983).
Weaver, R. F. & Weissmann, C. Nucleic Acids Res. 7, 1175–1193 (1979).
Bodary, S. & Mach, B. EMBO J. 1, 719–724 (1982).
Pech, M. et al. J. molec. Biol. (in the press).
Rechavi, G., Ram, D., Glazer, L., Zakut, R. & Givol, D. Proc. natn. Acad. Sci. U.S.A. 80, 855–859 (1983).
Nishioka, Y. & Leder, P. J. biol. Chem. 255, 3691–3694 (1980).
Bentley, D. L. & Rabbitts, T. H. Cell 32, 181–189 (1983).
Hozumi, N. et al. Proc. natn. Acad. Sci. U.S.A. 78, 7019–7023 (1981).
Max, E. E., Maizel, J. V. & Leder, P. J. biol. Chem. 256, 5116–5120 (1981).
Hieter, P. A., Max, E. E., Seidman, J. G., Maizel, J. V. & Leder, P. Cell 22, 197–207 (1980).
Early, P. et al. Cell 20, 313–319 (1980).
Ollo, R., Auffray, C., Morchamps, C. & Rougeon, F. Proc. natn. Acad. Sci. U.S.A. 78, 2442–2446 (1981).
Sakano, H., Maki, R., Kurosawa, Y., Roeder, W. & Tonegawa, S. Nature 286, 676–683 (1980).
Kataoka, T., Nikaido, T., Miyata, T., Moriwaki, K. & Honjo, T. J. biol. Chem. 257, 277–285 (1982).
Givol, D. et al. Nature 292, 426–430 (1981).
Seidman, J.G. & Leder, P. Nature 286, 779–783 (1980).
Sutcliffe, J. G. Cold Spring Harb. Symp. quant. Biol. 43, 77–90 (1979).
Seif, I., Khoury, G. & Dhar, R. Cell 18, 963–977 (1979).
Ullrich, A., Dull, T. J., Gray, A., Brosius, J. & Sures, I. Science 209, 612–615 (1980).
Robertson, M. A. et al. Nature 278, 370–372 (1979).
Birnstiel, M. et al. Proc. Alfred Benzon Symp. 13, 117–132 (1979).
Gebhard, W. & Zachau, H. G. J. molec. Biol. 170, 255–270 (1983).
Lawn, R. M., Efstratiadis, A., O'Connell, C. & Maniatis, T. Cell 21, 647–651 (1980).
Baralle, F. E., Shoulders, C. C. & Proudfoot, N. J. Cell 2l, 621–626 (1980).
Hyldig-Nielsen, J. J. et al. Nucleic Acids Res. 11, 5055–5071 (1983).
Mathis, D. J., Benoist, C. O., Williams, V. E. II, Kanter, M. R. & McDevitt, H. O. Cell 32, 745–754 (1983).
Malissen, M., Hunkapiller, T. & Hood, L. Science 221, 750–754 (1983).
Das, H. K., Lawrance, S. K. & Weissman, S. M. Proc. natn. Acad. Sci. U.S.A. 80, 3543–3547 (1983).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Falkner, F., Zachau, H. Correct transcription of an immunoglobulin κ gene requires an upstream fragment containing conserved sequence elements. Nature 310, 71–74 (1984). https://doi.org/10.1038/310071a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/310071a0
This article is cited by
-
Immunoglobulin germline gene variation and its impact on human disease
Genes & Immunity (2021)
-
Biological importance of OCT transcription factors in reprogramming and development
Experimental & Molecular Medicine (2021)
-
Immunoglobulin light chain (IGL) genes in torafugu: Genomic organization and identification of a third teleost IGL isotype
Scientific Reports (2017)
-
Characterization of the torafugu (Takifugu rubripes) immunoglobulin heavy chain gene locus
Immunogenetics (2015)
-
Alternative Promoters and Tissue-Specific Regulation of Mouse oct-1 Gene Transcription
Molecular Biology (2005)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.