Abstract
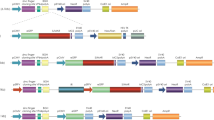
Genes lacking the intervening sequences that are present in otherwise apparently homologous genes have been found in some highly oncogenic retroviruses (some viral oncogenes) and in the normal cell genome (cDNA genes, pseudogenes or processed genes)1–10. Such cDNA genes might have arisen by reverse transcription of the spliced (intervening sequences removed) RNA transcript of the homologous genes. Another possible way for genes with intervening sequences to lose such sequences would be during replication in a retrovirus vector. To test this hypothesis we inserted genomic mouse α-globin DNA containing two intervening sequences11 into the DNA of an infectious retrovirus vector12 and report here that the intervening sequences were removed from DNA of progeny virus during virus replication.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Parker, R. C., Varmus, H. E. & Bishop, J. M. Proc. natn. Acad. Sci. U.S.A. 78, 5842–5846 (1981).
Goff, S. P., Gilboa, E., Witte, O. N. & Baltimore, D. Cell 22, 777–785 (1980).
Favera, R. D., Gelmann, E. P., Gallo, R. C. & Wong-Staal, F. Nature 292, 31–35 (1981).
Vennström, B. & Bishop, J. M. Cell 28, 135–143 (1982).
Chen, I. S. Y., Wilhelmsen, K. & Temin, H. M. J. Virol. (in the press).
Vanin, E. F., Goldberg, G. I., Tucker, P. W. & Smithies, O. Nature 286, 222–226 (1980).
Nishioka, Y. & Leder, A. Proc. natn. Acad. Sci. U.S.A. 77, 2806–2809 (1980).
Hollis, G. F., Hieter, P. A., McBride, D. W., Swan, D. & Leder, P. Nature 296, 321–325 (1982).
Sharp, P. A. 35th Ann. Symp. Fundamental Cancer Research (Raven, New York, in the press).
Wilde, C. D., Crowther, C. E., Gripe, T. P., Lee, M. G. S. & Cowan, N. J. Nature 297, 83–84 (1982).
Nishioka, Y. & Leder, P. Cell 18, 875–882 (1979).
Shimotohno, K. & Temin, H. M. Cell 26, 67–77 (1981).
O'Rear, J. J. & Temin, H. M. Proc. natn. Acad. Sci. U.S.A. 79, 1230–1234 (1982).
Watanabe, S. & Temin, H. M. Proc. natn. Acad. Sci. U.S.A. (in the press).
Ross, J. J. molec. Biol. 106, 403–420 (1976).
Varmus, H. & Swanstrom, R. in RNA Tumor Viruses, Molecular Biology of Tumor Viruses Pt 3 (eds Weiss, R. A. etai) 369–512 (Cold Spring Harbor Laboratory, New York, 1982).
Temin, H. M. Cell 27, 1–3 (1981).
Temin, H. M. Cell 28, 3–5 (1982).
Kinniburgh, A. J. & Ross, J. Cell 17, 915–921 (1979).
Temin, H. M. J. cell. Biochem. (in the press).
Tabak, H. F. & Flavell, R. A. Nucleic Acids Res. 5, 2321–2332 (1978).
Hirt, B. J. molec. Biol. 26, 365–369 (1967).
Chen, I. S. Y. & Temin, H. M. J. Virol. 41, 183–191 (1982).
Weller, S. K. & Temin, H. M. J. Virol. 39, 713–721 (1981).
Williams, B. G. & Blattner, F. R. J. Virol. 29, 555–575 (1979).
Maxam, A. M. & Gilbert, W. Meth. Enzym. 65, 499–560 (1980).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Shimotohno, K., Temin, H. Loss of intervening sequences in genomic mouse α-globin DNA inserted in an infectious retrovirus vector. Nature 299, 265–268 (1982). https://doi.org/10.1038/299265a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/299265a0
This article is cited by
-
The KT Jeang Retrovirology prize 2017: Michael Emerman
Retrovirology (2017)
-
Intratumoral gene delivery of anti-cathepsin L single-chain variable fragment by lentiviral vector inhibits tumor progression induced by human melanoma cells
Cancer Gene Therapy (2008)
-
Manufacture of a functional cDNA for the H-2Db molecule using a retroviral shuttle vector
Immunogenetics (1991)
-
Construction of chimaeric processed immunoglobulin genes containing mouse variable and human constant region sequences
Nature (1985)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.