Abstract
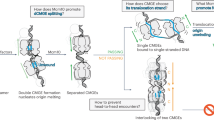
COLICINOGENIC plasmid El (ColEl) can be isolated from Escherichia coli as a supercoiled DNA molecule of 2.14 µm long or as a supercoiled DNA–protein complex1. Certain protein denaturing agents will convert this complex to an open circular (relaxed) DNA form with one single-strand break in the heavy (polyUG binding) strand of the duplex ColEl DNA (refs 2, 3). The induced single-strand break in ColEl DNA is located approximately 19% of the length of ColEl DNA from the single EcoRI restriction endonuclease site in the plasmid4. Analysis of ColEl replicating intermediates has shown that replication proceeds unidirectionally from an origin located near the position of the relaxation complex nick, approximately 19% from the single EcoRI restriction site 5–7. The site of the relaxation-complex nick has been located approximately 300 nucleotides away from the origin of ColEl DNA replication9. The nucleotide sequence of the origin region of ColEl DNA8,9 and of a small ColEl-type plasmid10 have been determined. A model for the possible role of the relaxation complex in ColEl DNA replication and/or conjugal transfer has been presented11. A major feature of this model is that the relaxation complex proteins may help promote association of plasmid DNA with a replicator and/or conjugal transfer site on the bacterial membrane. We describe here an electron microscopic analysis of EcoRI-cleaved ColEl and mini-ColEl DNA that were purified as stable DNA–membrane complexes from E. coli minicells. Evidence is presented that membrane-like structures attach to ColEl DNA and mini-ColEl DNA in a region that includes both the site of the origin/terminus of DNA replication and the site of single-strand cleavage by the relaxation complex.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Clewell, D. B. & Helinski, D. R. Proc. natn. Acad. Sci. U.S.A. 62, 1159–1166 (1969).
Clewell, D. B. & Helinski, D. R. Biochemistry 9, 4428–4440 (1970).
Blair, D. G., Clewell, D. B., Sheratt, D. J. & Helinski, D. R. Proc. natn. Acad. Sci. U.S.A. 68, 210–214 (1971).
Lovett, M. A., Guiney, D. & Helinski, D. R. Proc. natn. Acad. Sci. U.S.A. 71, 3854–3857 (1974).
Lovett, M. A., Katz, L. & Helinski, D. R. Nature 251, 337–340 (1974).
Inselburg, J. Proc. natn. Acad. Sci. U.S. A. 71, 2256–2259 (1974).
Tomizawa, J. I., Sakakibara, Y. & Kakefuda, T. Proc. natn. Acad. Sci. U.S.A. 71, 2260–2264 (1974).
Tomizawa, J., Ohmori, H. & Bird, R. Proc. natn. Acad. Sci. U.S.A. 74, 1865 (1977).
Bastia, D. Nucleic Acids Res. 4, 3123 (1977).
Bolivar, R. et al. Proc. natn. Acad. Sci. U.S.A. 74, 5265–5269 (1977).
Helinski, D. R. et al. in Cellular Modification and Genetic Transformation by Exogenous Nucleic Acids (eds Beers, R. F. & Tilghman, R. C.) 15–34 (1973).
Sueoka, N. & Hammers, J. Proc. natn. Acad. Sci. U.S.A. 71, 4787 (1974).
Hershfield, V., Boyer, H. W., Chow, L. & Helinski, D. R. J. Bact. 126, 447–453 (1976).
Osborn, M. J., Gander, J. E., Parisi, E. & Carson, J. J. biol. Chem. 247, 3962 (1972).
Alberts, B. M., Amodio, F. J., Jenkins, M., Girtmann, E. D. & Ferris, F. L. Cold Spring Harb. Symp. quant. Biol. 33, 289–305 (1968).
Davis, R. W., Simon, M. & Davidson, N. Meth. Enzym. 21, part D, 413–428 (1971).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
SPARKS, R., HELINSKI, D. Association of cellular membrane of E. coli minicells with the origins/terminus region of replication of plasmid ColEl DNA. Nature 277, 572–575 (1979). https://doi.org/10.1038/277572a0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/277572a0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.